meta-int-snw-23433
Interacting subnets
| Subnetwork | Dataset | Score | p-value 1 | p-value 2 | p-value 3 | Size | Highlighted genes |
|---|---|---|---|---|---|---|---|
| reg-snw-60 | tai-screen-luciferase | 4.243 | 1.88e-19 | 3.43e-03 | 4.71e-03 | 12 | 5 |
| int-snw-23433 | tai-screen-luciferase | 6.337 | 2.49e-134 | 4.62e-07 | 2.89e-03 | 10 | 7 |
| reg-snw-1121 | tai-screen-luciferase | 4.707 | 1.91e-24 | 5.19e-04 | 8.44e-04 | 5 | 3 |
| reg-snw-821 | tai-screen-luciferase | 4.344 | 1.76e-20 | 2.34e-03 | 3.35e-03 | 11 | 4 |
| reg-snw-51164 | tai-screen-luciferase | 4.586 | 4.41e-23 | 8.78e-04 | 1.37e-03 | 6 | 4 |
| reg-snw-57120 | tai-screen-luciferase | 4.450 | 1.33e-21 | 1.54e-03 | 2.29e-03 | 12 | 6 |
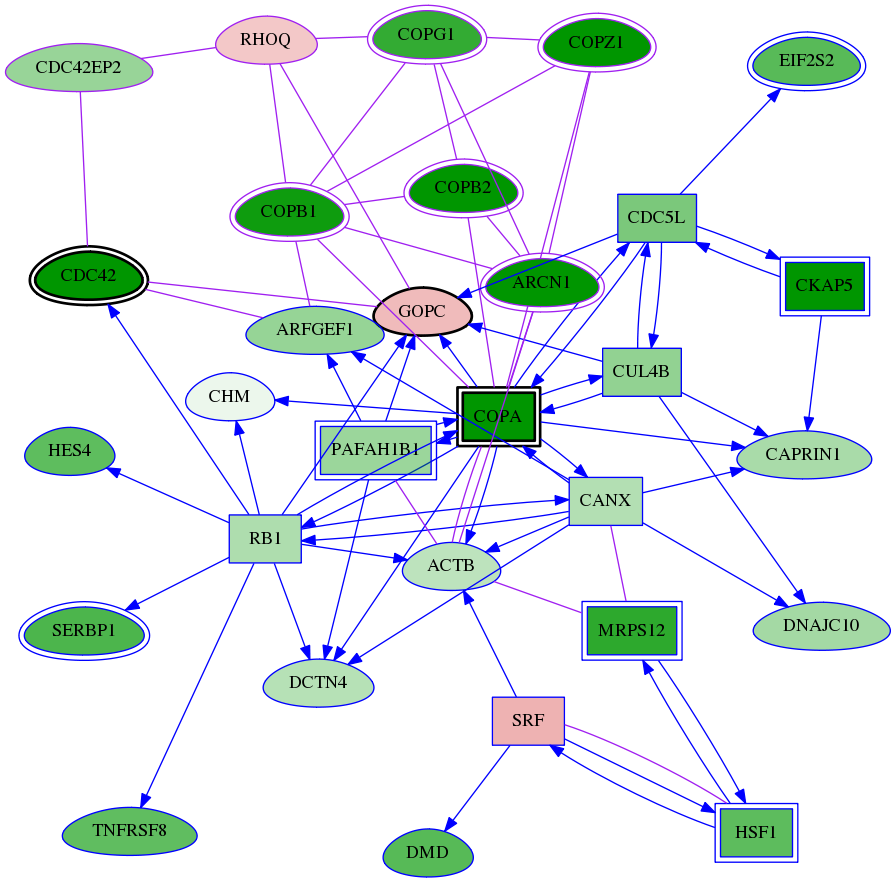
Genes (30)
| Gene Symbol | Entrez Gene ID | Frequency | tai-screen-luciferase gene score | Best subnetwork score | Degree | List-Gonzales GI | Tai-Hits |
|---|---|---|---|---|---|---|---|
| HES4 | 57801 | 19 | -4.153 | 4.450 | 10 | - | - |
| RHOQ | 23433 | 6 | 1.414 | 6.337 | 19 | - | - |
| DMD | 1756 | 15 | -4.338 | 4.243 | 53 | - | - |
| CDC42EP2 | 10435 | 6 | -2.649 | 6.337 | 5 | - | - |
| COPA | 1314 | 48 | -9.395 | 5.672 | 340 | Yes | Yes |
| PAFAH1B1 | 5048 | 18 | -2.583 | 4.586 | 55 | Yes | - |
| GOPC | 57120 | 19 | 1.749 | 4.450 | 68 | - | - |
| CAPRIN1 | 4076 | 14 | -2.226 | 4.688 | 42 | - | - |
| TNFRSF8 | 943 | 19 | -4.106 | 4.450 | 15 | - | - |
| CUL4B | 8450 | 21 | -2.809 | 5.269 | 142 | - | - |
| HSF1 | 3297 | 46 | -4.179 | 5.027 | 209 | - | Yes |
| ARFGEF1 | 10565 | 13 | -2.722 | 4.344 | 15 | - | - |
| CANX | 821 | 17 | -1.959 | 4.504 | 65 | - | - |
| COPB2 | 9276 | 48 | -13.168 | 9.063 | 41 | Yes | Yes |
| CKAP5 | 9793 | 46 | -7.214 | 5.672 | 130 | Yes | Yes |
| DNAJC10 | 54431 | 13 | -2.349 | 4.344 | 11 | - | - |
| COPZ1 | 22818 | 48 | -8.301 | 9.063 | 13 | Yes | Yes |
| RB1 | 5925 | 21 | -2.085 | 4.707 | 351 | - | - |
| EIF2S2 | 8894 | 31 | -4.320 | 5.672 | 103 | Yes | - |
| CHM | 1121 | 13 | -0.483 | 4.707 | 2 | - | - |
| CDC42 | 998 | 44 | -6.960 | 4.707 | 276 | Yes | Yes |
| DCTN4 | 51164 | 13 | -1.882 | 4.586 | 19 | - | - |
| MRPS12 | 6183 | 35 | -5.421 | 5.516 | 341 | Yes | - |
| SRF | 6722 | 15 | 1.994 | 4.243 | 23 | - | - |
| ACTB | 60 | 15 | -1.716 | 4.243 | 23 | - | - |
| ARCN1 | 372 | 48 | -8.232 | 9.063 | 118 | Yes | Yes |
| COPG1 | 22820 | 12 | -5.279 | 7.138 | 37 | Yes | Yes |
| COPB1 | 1315 | 39 | -6.221 | 9.063 | 118 | Yes | Yes |
| SERBP1 | 26135 | 35 | -4.612 | 5.516 | 106 | - | Yes |
| CDC5L | 988 | 34 | -3.419 | 5.672 | 155 | - | - |
Interactions (64)
| Gene Symbol 1 | Entrez Gene ID 1 | Gene Symbol 2 | Entrez Gene ID 2 | Type | Direction | Origin databases / Sources |
|---|---|---|---|---|---|---|
| TNFRSF8 | 943 | RB1 | 5925 | pd | < | reg.pazar.txt: no annot |
| ARCN1 | 372 | COPZ1 | 22818 | pp | -- | int.I2D: BioGrid_Yeast, IntAct_Yeast, MINT_Yeast, Tarassov_PCA, YeastMedium, INTEROLOG |
| PAFAH1B1 | 5048 | ARFGEF1 | 10565 | pd | > | reg.ITFP.txt: no annot |
| COPZ1 | 22818 | COPG1 | 22820 | pp | -- | int.I2D: BioGrid, HPRD, BIND; int.HPRD: in vitro, in vivo, yeast 2-hybrid; int.DIP: MI:0915(physical association) |
| ACTB | 60 | CANX | 821 | pd | < | reg.ITFP.txt: no annot |
| COPA | 1314 | RB1 | 5925 | pd | <> | reg.ITFP.txt: no annot |
| HSF1 | 3297 | SRF | 6722 | pd | <> | reg.ITFP.txt: no annot |
| HSF1 | 3297 | SRF | 6722 | pp | -- | int.Yu: muscle |
| COPA | 1314 | DCTN4 | 51164 | pd | > | reg.ITFP.txt: no annot |
| CDC5L | 988 | CUL4B | 8450 | pd | <> | reg.ITFP.txt: no annot |
| COPB2 | 9276 | COPG1 | 22820 | pp | -- | int.I2D: BioGrid |
| RB1 | 5925 | GOPC | 57120 | pd | > | reg.ITFP.txt: no annot |
| CANX | 821 | ARFGEF1 | 10565 | pd | > | reg.ITFP.txt: no annot |
| CANX | 821 | CAPRIN1 | 4076 | pd | > | reg.ITFP.txt: no annot |
| COPB1 | 1315 | RHOQ | 23433 | pp | -- | int.I2D: HPRD, BCI; int.HPRD: in vivo |
| RB1 | 5925 | HES4 | 57801 | pd | > | reg.pazar.txt: no annot |
| CDC42EP2 | 10435 | RHOQ | 23433 | pp | -- | int.I2D: BioGrid, HPRD, BIND; int.HPRD: in vitro, yeast 2-hybrid |
| CHM | 1121 | COPA | 1314 | pd | < | reg.ITFP.txt: no annot |
| RB1 | 5925 | SERBP1 | 26135 | pd | > | reg.pazar.txt: no annot |
| ACTB | 60 | MRPS12 | 6183 | pp | -- | int.I2D: BioGrid_Yeast |
| ARCN1 | 372 | COPG1 | 22820 | pp | -- | int.I2D: HPRD; int.HPRD: in vivo |
| COPG1 | 22820 | RHOQ | 23433 | pp | -- | int.I2D: HPRD; int.HPRD: in vivo |
| CUL4B | 8450 | GOPC | 57120 | pd | > | reg.ITFP.txt: no annot |
| PAFAH1B1 | 5048 | GOPC | 57120 | pd | > | reg.ITFP.txt: no annot |
| CAPRIN1 | 4076 | CKAP5 | 9793 | pd | < | reg.ITFP.txt: no annot |
| COPB1 | 1315 | COPG1 | 22820 | pp | -- | int.I2D: BioGrid, HPRD; int.HPRD: in vitro, in vivo, yeast 2-hybrid; int.DIP: MI:0915(physical association) |
| CDC42 | 998 | ARFGEF1 | 10565 | pp | -- | int.I2D: IntAct, SOURAV_MAPK_LOW |
| RHOQ | 23433 | GOPC | 57120 | pp | -- | int.I2D: BioGrid, HPRD; int.HPRD: in vitro, in vivo, yeast 2-hybrid |
| ACTB | 60 | RB1 | 5925 | pd | < | reg.ITFP.txt: no annot |
| ACTB | 60 | COPA | 1314 | pd | < | reg.ITFP.txt: no annot |
| ACTB | 60 | COPA | 1314 | pp | -- | int.I2D: BioGrid_Yeast |
| CAPRIN1 | 4076 | CUL4B | 8450 | pd | < | reg.ITFP.txt: no annot |
| COPA | 1314 | COPB1 | 1315 | pp | -- | int.I2D: BioGrid, BioGrid_Yeast, BIND_Yeast, IntAct_Yeast, MINT_Yeast, Krogan_Core, YeastHigh |
| CDC5L | 988 | CKAP5 | 9793 | pd | <> | reg.ITFP.txt: no annot |
| CDC42 | 998 | CDC42EP2 | 10435 | pp | -- | int.Intact: MI:0915(physical association); int.I2D: BioGrid, HPRD, BIND, IntAct, SOURAV_MAPK_HIGH; int.HPRD: in vitro, in vivo |
| PAFAH1B1 | 5048 | DCTN4 | 51164 | pd | > | reg.ITFP.txt: no annot |
| DMD | 1756 | SRF | 6722 | pd | < | reg.pazar.txt: no annot; reg.TRANSFAC.txt: no annot |
| CANX | 821 | RB1 | 5925 | pd | <> | reg.ITFP.txt: no annot |
| ACTB | 60 | PAFAH1B1 | 5048 | pp | -- | int.I2D: BioGrid_Yeast |
| COPA | 1314 | COPZ1 | 22818 | pp | -- | int.I2D: BioGrid_Yeast, IntAct_Yeast, Krogan_Core, MINT_Yeast, YeastMedium |
| CANX | 821 | MRPS12 | 6183 | pp | -- | int.I2D: BioGrid_Yeast |
| CDC42 | 998 | GOPC | 57120 | pp | -- | int.I2D: BioGrid |
| COPA | 1314 | COPB2 | 9276 | pp | -- | int.I2D: BioGrid, HPRD, BCI; int.HPRD: in vivo |
| COPB1 | 1315 | COPZ1 | 22818 | pp | -- | int.I2D: BioGrid_Yeast, IntAct_Yeast, Krogan_Core, MINT_Yeast, Tarassov_PCA, YeastMedium; int.DIP: MI:0915(physical association) |
| CANX | 821 | COPA | 1314 | pd | <> | reg.ITFP.txt: no annot |
| COPA | 1314 | CAPRIN1 | 4076 | pd | > | reg.ITFP.txt: no annot |
| ACTB | 60 | ARCN1 | 372 | pp | -- | int.I2D: BioGrid_Yeast |
| CUL4B | 8450 | DNAJC10 | 54431 | pd | > | reg.ITFP.txt: no annot |
| HSF1 | 3297 | MRPS12 | 6183 | pd | <> | reg.ITFP.txt: no annot |
| RB1 | 5925 | DCTN4 | 51164 | pd | > | reg.ITFP.txt: no annot |
| COPA | 1314 | GOPC | 57120 | pd | > | reg.ITFP.txt: no annot |
| CDC5L | 988 | GOPC | 57120 | pd | > | reg.ITFP.txt: no annot |
| CDC5L | 988 | COPA | 1314 | pd | <> | reg.ITFP.txt: no annot |
| COPA | 1314 | CUL4B | 8450 | pd | <> | reg.ITFP.txt: no annot |
| CDC5L | 988 | EIF2S2 | 8894 | pd | > | reg.ITFP.txt: no annot |
| COPB1 | 1315 | COPB2 | 9276 | pp | -- | int.I2D: BioGrid |
| CHM | 1121 | RB1 | 5925 | pd | < | reg.ITFP.txt: no annot |
| ARCN1 | 372 | COPB1 | 1315 | pp | -- | int.I2D: BCI, BioGrid, BioGrid_Yeast, IntAct_Yeast, INTEROLOG, MINT_Yeast, YeastHigh, BIND, BIND_Yeast, HPRD, Krogan_Core, MIPS; int.HPRD: in vivo |
| CANX | 821 | DCTN4 | 51164 | pd | > | reg.ITFP.txt: no annot |
| ARCN1 | 372 | COPA | 1314 | pp | -- | int.I2D: BCI, BioGrid_Yeast, IntAct_Yeast, MINT_Yeast, YeastHigh, HPRD, Krogan_Core; int.HPRD: in vivo |
| CDC42 | 998 | RB1 | 5925 | pd | < | reg.pazar.txt: no annot |
| CANX | 821 | DNAJC10 | 54431 | pd | > | reg.ITFP.txt: no annot |
| COPB1 | 1315 | ARFGEF1 | 10565 | pp | -- | int.I2D: BioGrid_Yeast |
| ARCN1 | 372 | COPB2 | 9276 | pp | -- | int.I2D: HPRD, BCI; int.HPRD: in vivo |
| ACTB | 60 | SRF | 6722 | pd | < | reg.pazar.txt: no annot; reg.oreganno.txt: no annot |
| COPA | 1314 | PAFAH1B1 | 5048 | pd | <> | reg.ITFP.txt: no annot |
Related GO terms (524)
| Accession number | Name | Hypergeometric test | Corrected p-value | Enrichment ratio | Occurrence in subnetwork | Occurrences in all snw genes | Occurrences in all int/reg genes |
|---|---|---|---|---|---|---|---|
| GO:0048205 | COPI coating of Golgi vesicle | 3.85e-14 | 6.28e-10 | 7.972 | 6 | 6 | 13 |
| GO:0030126 | COPI vesicle coat | 3.85e-14 | 6.28e-10 | 7.972 | 6 | 6 | 13 |
| GO:0006890 | retrograde vesicle-mediated transport, Golgi to ER | 3.91e-12 | 6.39e-08 | 7.029 | 6 | 6 | 25 |
| GO:0061024 | membrane organization | 6.80e-09 | 1.11e-04 | 4.705 | 7 | 11 | 146 |
| GO:0005829 | cytosol | 3.09e-07 | 5.05e-03 | 1.852 | 17 | 74 | 2562 |
| GO:0006886 | intracellular protein transport | 6.25e-07 | 1.02e-02 | 4.238 | 6 | 6 | 173 |
| GO:0006891 | intra-Golgi vesicle-mediated transport | 3.75e-06 | 6.11e-02 | 6.585 | 3 | 3 | 17 |
| GO:0016020 | membrane | 4.77e-06 | 7.78e-02 | 2.018 | 13 | 48 | 1746 |
| GO:0051683 | establishment of Golgi localization | 9.79e-06 | 1.60e-01 | 8.503 | 2 | 2 | 3 |
| GO:0043234 | protein complex | 1.50e-05 | 2.45e-01 | 3.444 | 6 | 9 | 300 |
| GO:0072384 | organelle transport along microtubule | 1.96e-05 | 3.19e-01 | 8.088 | 2 | 2 | 4 |
| GO:0007097 | nuclear migration | 6.82e-05 | 1.00e+00 | 7.280 | 2 | 2 | 7 |
| GO:0090136 | epithelial cell-cell adhesion | 1.17e-04 | 1.00e+00 | 6.918 | 2 | 2 | 9 |
| GO:0031274 | positive regulation of pseudopodium assembly | 1.46e-04 | 1.00e+00 | 6.766 | 2 | 2 | 10 |
| GO:0005198 | structural molecule activity | 1.95e-04 | 1.00e+00 | 3.775 | 4 | 5 | 159 |
| GO:0050998 | nitric-oxide synthase binding | 3.86e-04 | 1.00e+00 | 6.088 | 2 | 3 | 16 |
| GO:0017022 | myosin binding | 4.91e-04 | 1.00e+00 | 5.918 | 2 | 2 | 18 |
| GO:0005737 | cytoplasm | 5.95e-04 | 1.00e+00 | 1.130 | 16 | 65 | 3976 |
| GO:0009306 | protein secretion | 6.71e-04 | 1.00e+00 | 5.695 | 2 | 2 | 21 |
| GO:0036464 | cytoplasmic ribonucleoprotein granule | 7.37e-04 | 1.00e+00 | 5.628 | 2 | 5 | 22 |
| GO:0051491 | positive regulation of filopodium assembly | 8.07e-04 | 1.00e+00 | 5.564 | 2 | 2 | 23 |
| GO:0000139 | Golgi membrane | 8.10e-04 | 1.00e+00 | 2.737 | 5 | 7 | 408 |
| GO:0005515 | protein binding | 1.14e-03 | 1.00e+00 | 0.829 | 20 | 87 | 6127 |
| GO:0051219 | phosphoprotein binding | 1.57e-03 | 1.00e+00 | 5.088 | 2 | 4 | 32 |
| GO:0030036 | actin cytoskeleton organization | 1.68e-03 | 1.00e+00 | 3.661 | 3 | 3 | 129 |
| GO:0030038 | contractile actin filament bundle assembly | 1.84e-03 | 1.00e+00 | 9.088 | 1 | 1 | 1 |
| GO:0045556 | positive regulation of TRAIL biosynthetic process | 1.84e-03 | 1.00e+00 | 9.088 | 1 | 1 | 1 |
| GO:0090287 | regulation of cellular response to growth factor stimulus | 1.84e-03 | 1.00e+00 | 9.088 | 1 | 1 | 1 |
| GO:0000235 | astral microtubule | 1.84e-03 | 1.00e+00 | 9.088 | 1 | 1 | 1 |
| GO:0090284 | positive regulation of protein glycosylation in Golgi | 1.84e-03 | 1.00e+00 | 9.088 | 1 | 1 | 1 |
| GO:0090230 | regulation of centromere complex assembly | 1.84e-03 | 1.00e+00 | 9.088 | 1 | 1 | 1 |
| GO:0032427 | GBD domain binding | 1.84e-03 | 1.00e+00 | 9.088 | 1 | 1 | 1 |
| GO:0043004 | cytoplasmic sequestering of CFTR protein | 1.84e-03 | 1.00e+00 | 9.088 | 1 | 1 | 1 |
| GO:0021691 | cerebellar Purkinje cell layer maturation | 1.84e-03 | 1.00e+00 | 9.088 | 1 | 1 | 1 |
| GO:0051660 | establishment of centrosome localization | 1.84e-03 | 1.00e+00 | 9.088 | 1 | 1 | 1 |
| GO:0034975 | protein folding in endoplasmic reticulum | 1.84e-03 | 1.00e+00 | 9.088 | 1 | 1 | 1 |
| GO:0046016 | positive regulation of transcription by glucose | 1.84e-03 | 1.00e+00 | 9.088 | 1 | 1 | 1 |
| GO:0046469 | platelet activating factor metabolic process | 1.84e-03 | 1.00e+00 | 9.088 | 1 | 1 | 1 |
| GO:0034332 | adherens junction organization | 1.98e-03 | 1.00e+00 | 4.918 | 2 | 4 | 36 |
| GO:0021766 | hippocampus development | 2.32e-03 | 1.00e+00 | 4.802 | 2 | 3 | 39 |
| GO:0044822 | poly(A) RNA binding | 2.86e-03 | 1.00e+00 | 1.821 | 7 | 42 | 1078 |
| GO:0034613 | cellular protein localization | 2.95e-03 | 1.00e+00 | 4.628 | 2 | 2 | 44 |
| GO:0021762 | substantia nigra development | 3.21e-03 | 1.00e+00 | 4.564 | 2 | 2 | 46 |
| GO:0008344 | adult locomotory behavior | 3.35e-03 | 1.00e+00 | 4.533 | 2 | 2 | 47 |
| GO:0000978 | RNA polymerase II core promoter proximal region sequence-specific DNA binding | 3.55e-03 | 1.00e+00 | 3.280 | 3 | 4 | 168 |
| GO:0010256 | endomembrane system organization | 3.67e-03 | 1.00e+00 | 8.088 | 1 | 1 | 2 |
| GO:0071987 | WD40-repeat domain binding | 3.67e-03 | 1.00e+00 | 8.088 | 1 | 1 | 2 |
| GO:0060661 | submandibular salivary gland formation | 3.67e-03 | 1.00e+00 | 8.088 | 1 | 1 | 2 |
| GO:0036035 | osteoclast development | 3.67e-03 | 1.00e+00 | 8.088 | 1 | 1 | 2 |
| GO:0014819 | regulation of skeletal muscle contraction | 3.67e-03 | 1.00e+00 | 8.088 | 1 | 1 | 2 |
| GO:0086001 | cardiac muscle cell action potential | 3.67e-03 | 1.00e+00 | 8.088 | 1 | 1 | 2 |
| GO:0071338 | positive regulation of hair follicle cell proliferation | 3.67e-03 | 1.00e+00 | 8.088 | 1 | 1 | 2 |
| GO:0051081 | nuclear envelope disassembly | 3.67e-03 | 1.00e+00 | 8.088 | 1 | 1 | 2 |
| GO:0002176 | male germ cell proliferation | 3.67e-03 | 1.00e+00 | 8.088 | 1 | 1 | 2 |
| GO:0072422 | signal transduction involved in DNA damage checkpoint | 3.67e-03 | 1.00e+00 | 8.088 | 1 | 1 | 2 |
| GO:0090135 | actin filament branching | 3.67e-03 | 1.00e+00 | 8.088 | 1 | 1 | 2 |
| GO:0031134 | sister chromatid biorientation | 3.67e-03 | 1.00e+00 | 8.088 | 1 | 1 | 2 |
| GO:0010736 | serum response element binding | 3.67e-03 | 1.00e+00 | 8.088 | 1 | 1 | 2 |
| GO:0007070 | negative regulation of transcription from RNA polymerase II promoter during mitosis | 3.67e-03 | 1.00e+00 | 8.088 | 1 | 1 | 2 |
| GO:0007030 | Golgi organization | 3.79e-03 | 1.00e+00 | 4.444 | 2 | 3 | 50 |
| GO:0030424 | axon | 3.79e-03 | 1.00e+00 | 3.246 | 3 | 4 | 172 |
| GO:0030175 | filopodium | 4.25e-03 | 1.00e+00 | 4.360 | 2 | 2 | 53 |
| GO:0001162 | RNA polymerase II intronic transcription regulatory region sequence-specific DNA binding | 5.50e-03 | 1.00e+00 | 7.503 | 1 | 1 | 3 |
| GO:0014809 | regulation of skeletal muscle contraction by regulation of release of sequestered calcium ion | 5.50e-03 | 1.00e+00 | 7.503 | 1 | 1 | 3 |
| GO:0060947 | cardiac vascular smooth muscle cell differentiation | 5.50e-03 | 1.00e+00 | 7.503 | 1 | 1 | 3 |
| GO:0003161 | cardiac conduction system development | 5.50e-03 | 1.00e+00 | 7.503 | 1 | 1 | 3 |
| GO:0071459 | protein localization to chromosome, centromeric region | 5.50e-03 | 1.00e+00 | 7.503 | 1 | 1 | 3 |
| GO:1900222 | negative regulation of beta-amyloid clearance | 5.50e-03 | 1.00e+00 | 7.503 | 1 | 1 | 3 |
| GO:1901385 | regulation of voltage-gated calcium channel activity | 5.50e-03 | 1.00e+00 | 7.503 | 1 | 1 | 3 |
| GO:0071930 | negative regulation of transcription involved in G1/S transition of mitotic cell cycle | 5.50e-03 | 1.00e+00 | 7.503 | 1 | 1 | 3 |
| GO:0045505 | dynein intermediate chain binding | 5.50e-03 | 1.00e+00 | 7.503 | 1 | 1 | 3 |
| GO:0022028 | tangential migration from the subventricular zone to the olfactory bulb | 5.50e-03 | 1.00e+00 | 7.503 | 1 | 1 | 3 |
| GO:0016671 | oxidoreductase activity, acting on a sulfur group of donors, disulfide as acceptor | 5.50e-03 | 1.00e+00 | 7.503 | 1 | 1 | 3 |
| GO:0030951 | establishment or maintenance of microtubule cytoskeleton polarity | 5.50e-03 | 1.00e+00 | 7.503 | 1 | 1 | 3 |
| GO:0033561 | regulation of water loss via skin | 5.50e-03 | 1.00e+00 | 7.503 | 1 | 1 | 3 |
| GO:0005850 | eukaryotic translation initiation factor 2 complex | 5.50e-03 | 1.00e+00 | 7.503 | 1 | 1 | 3 |
| GO:0034663 | endoplasmic reticulum chaperone complex | 5.50e-03 | 1.00e+00 | 7.503 | 1 | 1 | 3 |
| GO:1902083 | negative regulation of peptidyl-cysteine S-nitrosylation | 5.50e-03 | 1.00e+00 | 7.503 | 1 | 2 | 3 |
| GO:0021540 | corpus callosum morphogenesis | 5.50e-03 | 1.00e+00 | 7.503 | 1 | 1 | 3 |
| GO:0071922 | regulation of cohesin localization to chromatin | 7.33e-03 | 1.00e+00 | 7.088 | 1 | 2 | 4 |
| GO:0034191 | apolipoprotein A-I receptor binding | 7.33e-03 | 1.00e+00 | 7.088 | 1 | 1 | 4 |
| GO:0010669 | epithelial structure maintenance | 7.33e-03 | 1.00e+00 | 7.088 | 1 | 1 | 4 |
| GO:0060684 | epithelial-mesenchymal cell signaling | 7.33e-03 | 1.00e+00 | 7.088 | 1 | 1 | 4 |
| GO:0051835 | positive regulation of synapse structural plasticity | 7.33e-03 | 1.00e+00 | 7.088 | 1 | 1 | 4 |
| GO:0015036 | disulfide oxidoreductase activity | 7.33e-03 | 1.00e+00 | 7.088 | 1 | 1 | 4 |
| GO:0043550 | regulation of lipid kinase activity | 7.33e-03 | 1.00e+00 | 7.088 | 1 | 1 | 4 |
| GO:0016013 | syntrophin complex | 7.33e-03 | 1.00e+00 | 7.088 | 1 | 2 | 4 |
| GO:0090231 | regulation of spindle checkpoint | 7.33e-03 | 1.00e+00 | 7.088 | 1 | 1 | 4 |
| GO:0003257 | positive regulation of transcription from RNA polymerase II promoter involved in myocardial precursor cell differentiation | 7.33e-03 | 1.00e+00 | 7.088 | 1 | 1 | 4 |
| GO:0048664 | neuron fate determination | 7.33e-03 | 1.00e+00 | 7.088 | 1 | 1 | 4 |
| GO:0010735 | positive regulation of transcription via serum response element binding | 7.33e-03 | 1.00e+00 | 7.088 | 1 | 1 | 4 |
| GO:0005968 | Rab-protein geranylgeranyltransferase complex | 7.33e-03 | 1.00e+00 | 7.088 | 1 | 1 | 4 |
| GO:0035189 | Rb-E2F complex | 7.33e-03 | 1.00e+00 | 7.088 | 1 | 1 | 4 |
| GO:0034349 | glial cell apoptotic process | 7.33e-03 | 1.00e+00 | 7.088 | 1 | 1 | 4 |
| GO:0034088 | maintenance of mitotic sister chromatid cohesion | 7.33e-03 | 1.00e+00 | 7.088 | 1 | 1 | 4 |
| GO:0031175 | neuron projection development | 8.34e-03 | 1.00e+00 | 3.859 | 2 | 2 | 75 |
| GO:0038096 | Fc-gamma receptor signaling pathway involved in phagocytosis | 8.77e-03 | 1.00e+00 | 3.821 | 2 | 4 | 77 |
| GO:0036336 | dendritic cell migration | 9.16e-03 | 1.00e+00 | 6.766 | 1 | 1 | 5 |
| GO:2000651 | positive regulation of sodium ion transmembrane transporter activity | 9.16e-03 | 1.00e+00 | 6.766 | 1 | 1 | 5 |
| GO:0035088 | establishment or maintenance of apical/basal cell polarity | 9.16e-03 | 1.00e+00 | 6.766 | 1 | 1 | 5 |
| GO:0048667 | cell morphogenesis involved in neuron differentiation | 9.16e-03 | 1.00e+00 | 6.766 | 1 | 1 | 5 |
| GO:0031256 | leading edge membrane | 9.16e-03 | 1.00e+00 | 6.766 | 1 | 1 | 5 |
| GO:0044233 | ER-mitochondrion membrane contact site | 9.16e-03 | 1.00e+00 | 6.766 | 1 | 1 | 5 |
| GO:0035912 | dorsal aorta morphogenesis | 9.16e-03 | 1.00e+00 | 6.766 | 1 | 1 | 5 |
| GO:0001515 | opioid peptide activity | 9.16e-03 | 1.00e+00 | 6.766 | 1 | 1 | 5 |
| GO:0031465 | Cul4B-RING E3 ubiquitin ligase complex | 9.16e-03 | 1.00e+00 | 6.766 | 1 | 1 | 5 |
| GO:0043043 | peptide biosynthetic process | 9.16e-03 | 1.00e+00 | 6.766 | 1 | 1 | 5 |
| GO:0031023 | microtubule organizing center organization | 9.16e-03 | 1.00e+00 | 6.766 | 1 | 1 | 5 |
| GO:0004663 | Rab geranylgeranyltransferase activity | 9.16e-03 | 1.00e+00 | 6.766 | 1 | 1 | 5 |
| GO:0008134 | transcription factor binding | 1.02e-02 | 1.00e+00 | 2.730 | 3 | 5 | 246 |
| GO:0043353 | enucleate erythrocyte differentiation | 1.10e-02 | 1.00e+00 | 6.503 | 1 | 1 | 6 |
| GO:0007143 | female meiotic division | 1.10e-02 | 1.00e+00 | 6.503 | 1 | 1 | 6 |
| GO:0044458 | motile cilium assembly | 1.10e-02 | 1.00e+00 | 6.503 | 1 | 1 | 6 |
| GO:0051150 | regulation of smooth muscle cell differentiation | 1.10e-02 | 1.00e+00 | 6.503 | 1 | 1 | 6 |
| GO:0018344 | protein geranylgeranylation | 1.10e-02 | 1.00e+00 | 6.503 | 1 | 1 | 6 |
| GO:0060789 | hair follicle placode formation | 1.10e-02 | 1.00e+00 | 6.503 | 1 | 1 | 6 |
| GO:0008090 | retrograde axon cargo transport | 1.10e-02 | 1.00e+00 | 6.503 | 1 | 1 | 6 |
| GO:0048554 | positive regulation of metalloenzyme activity | 1.10e-02 | 1.00e+00 | 6.503 | 1 | 1 | 6 |
| GO:0045842 | positive regulation of mitotic metaphase/anaphase transition | 1.10e-02 | 1.00e+00 | 6.503 | 1 | 1 | 6 |
| GO:0001667 | ameboidal-type cell migration | 1.10e-02 | 1.00e+00 | 6.503 | 1 | 1 | 6 |
| GO:0000974 | Prp19 complex | 1.10e-02 | 1.00e+00 | 6.503 | 1 | 1 | 6 |
| GO:0030957 | Tat protein binding | 1.10e-02 | 1.00e+00 | 6.503 | 1 | 1 | 6 |
| GO:0034452 | dynactin binding | 1.10e-02 | 1.00e+00 | 6.503 | 1 | 1 | 6 |
| GO:0043025 | neuronal cell body | 1.11e-02 | 1.00e+00 | 2.684 | 3 | 5 | 254 |
| GO:0019886 | antigen processing and presentation of exogenous peptide antigen via MHC class II | 1.23e-02 | 1.00e+00 | 3.564 | 2 | 3 | 92 |
| GO:0005200 | structural constituent of cytoskeleton | 1.26e-02 | 1.00e+00 | 3.548 | 2 | 6 | 93 |
| GO:0060261 | positive regulation of transcription initiation from RNA polymerase II promoter | 1.28e-02 | 1.00e+00 | 6.280 | 1 | 1 | 7 |
| GO:0006893 | Golgi to plasma membrane transport | 1.28e-02 | 1.00e+00 | 6.280 | 1 | 1 | 7 |
| GO:0060055 | angiogenesis involved in wound healing | 1.28e-02 | 1.00e+00 | 6.280 | 1 | 1 | 7 |
| GO:0060136 | embryonic process involved in female pregnancy | 1.28e-02 | 1.00e+00 | 6.280 | 1 | 1 | 7 |
| GO:0043497 | regulation of protein heterodimerization activity | 1.28e-02 | 1.00e+00 | 6.280 | 1 | 1 | 7 |
| GO:0050658 | RNA transport | 1.28e-02 | 1.00e+00 | 6.280 | 1 | 1 | 7 |
| GO:0003334 | keratinocyte development | 1.28e-02 | 1.00e+00 | 6.280 | 1 | 1 | 7 |
| GO:0051988 | regulation of attachment of spindle microtubules to kinetochore | 1.28e-02 | 1.00e+00 | 6.280 | 1 | 1 | 7 |
| GO:0070914 | UV-damage excision repair | 1.28e-02 | 1.00e+00 | 6.280 | 1 | 1 | 7 |
| GO:0001961 | positive regulation of cytokine-mediated signaling pathway | 1.28e-02 | 1.00e+00 | 6.280 | 1 | 1 | 7 |
| GO:0002162 | dystroglycan binding | 1.28e-02 | 1.00e+00 | 6.280 | 1 | 1 | 7 |
| GO:0030157 | pancreatic juice secretion | 1.28e-02 | 1.00e+00 | 6.280 | 1 | 1 | 7 |
| GO:0017145 | stem cell division | 1.28e-02 | 1.00e+00 | 6.280 | 1 | 1 | 7 |
| GO:0000930 | gamma-tubulin complex | 1.28e-02 | 1.00e+00 | 6.280 | 1 | 1 | 7 |
| GO:0034259 | negative regulation of Rho GTPase activity | 1.28e-02 | 1.00e+00 | 6.280 | 1 | 1 | 7 |
| GO:0001764 | neuron migration | 1.34e-02 | 1.00e+00 | 3.503 | 2 | 2 | 96 |
| GO:0090005 | negative regulation of establishment of protein localization to plasma membrane | 1.46e-02 | 1.00e+00 | 6.088 | 1 | 1 | 8 |
| GO:0090009 | primitive streak formation | 1.46e-02 | 1.00e+00 | 6.088 | 1 | 1 | 8 |
| GO:0031512 | motile primary cilium | 1.46e-02 | 1.00e+00 | 6.088 | 1 | 1 | 8 |
| GO:0043589 | skin morphogenesis | 1.46e-02 | 1.00e+00 | 6.088 | 1 | 1 | 8 |
| GO:0007289 | spermatid nucleus differentiation | 1.46e-02 | 1.00e+00 | 6.088 | 1 | 1 | 8 |
| GO:0005522 | profilin binding | 1.46e-02 | 1.00e+00 | 6.088 | 1 | 1 | 8 |
| GO:0070688 | MLL5-L complex | 1.46e-02 | 1.00e+00 | 6.088 | 1 | 1 | 8 |
| GO:0005869 | dynactin complex | 1.46e-02 | 1.00e+00 | 6.088 | 1 | 1 | 8 |
| GO:0051489 | regulation of filopodium assembly | 1.46e-02 | 1.00e+00 | 6.088 | 1 | 1 | 8 |
| GO:0061003 | positive regulation of dendritic spine morphogenesis | 1.46e-02 | 1.00e+00 | 6.088 | 1 | 1 | 8 |
| GO:0008360 | regulation of cell shape | 1.61e-02 | 1.00e+00 | 3.360 | 2 | 2 | 106 |
| GO:0014069 | postsynaptic density | 1.61e-02 | 1.00e+00 | 3.360 | 2 | 3 | 106 |
| GO:0097284 | hepatocyte apoptotic process | 1.64e-02 | 1.00e+00 | 5.918 | 1 | 1 | 9 |
| GO:0000983 | RNA polymerase II core promoter sequence-specific DNA binding transcription factor activity | 1.64e-02 | 1.00e+00 | 5.918 | 1 | 1 | 9 |
| GO:0000075 | cell cycle checkpoint | 1.64e-02 | 1.00e+00 | 5.918 | 1 | 1 | 9 |
| GO:0021895 | cerebral cortex neuron differentiation | 1.64e-02 | 1.00e+00 | 5.918 | 1 | 1 | 9 |
| GO:0030837 | negative regulation of actin filament polymerization | 1.64e-02 | 1.00e+00 | 5.918 | 1 | 1 | 9 |
| GO:0032319 | regulation of Rho GTPase activity | 1.64e-02 | 1.00e+00 | 5.918 | 1 | 1 | 9 |
| GO:0045059 | positive thymic T cell selection | 1.64e-02 | 1.00e+00 | 5.918 | 1 | 1 | 9 |
| GO:0015630 | microtubule cytoskeleton | 1.79e-02 | 1.00e+00 | 3.280 | 2 | 4 | 112 |
| GO:0034237 | protein kinase A regulatory subunit binding | 1.82e-02 | 1.00e+00 | 5.766 | 1 | 1 | 10 |
| GO:0051787 | misfolded protein binding | 1.82e-02 | 1.00e+00 | 5.766 | 1 | 1 | 10 |
| GO:0005798 | Golgi-associated vesicle | 1.82e-02 | 1.00e+00 | 5.766 | 1 | 1 | 10 |
| GO:0042535 | positive regulation of tumor necrosis factor biosynthetic process | 1.82e-02 | 1.00e+00 | 5.766 | 1 | 1 | 10 |
| GO:0017049 | GTP-Rho binding | 1.82e-02 | 1.00e+00 | 5.766 | 1 | 1 | 10 |
| GO:0001076 | RNA polymerase II transcription factor binding transcription factor activity | 1.82e-02 | 1.00e+00 | 5.766 | 1 | 1 | 10 |
| GO:0090303 | positive regulation of wound healing | 1.82e-02 | 1.00e+00 | 5.766 | 1 | 1 | 10 |
| GO:0061029 | eyelid development in camera-type eye | 1.82e-02 | 1.00e+00 | 5.766 | 1 | 1 | 10 |
| GO:0001675 | acrosome assembly | 1.82e-02 | 1.00e+00 | 5.766 | 1 | 1 | 10 |
| GO:0060218 | hematopoietic stem cell differentiation | 1.82e-02 | 1.00e+00 | 5.766 | 1 | 1 | 10 |
| GO:0060047 | heart contraction | 1.82e-02 | 1.00e+00 | 5.766 | 1 | 1 | 10 |
| GO:0045502 | dynein binding | 2.00e-02 | 1.00e+00 | 5.628 | 1 | 1 | 11 |
| GO:0045176 | apical protein localization | 2.00e-02 | 1.00e+00 | 5.628 | 1 | 1 | 11 |
| GO:0017166 | vinculin binding | 2.00e-02 | 1.00e+00 | 5.628 | 1 | 1 | 11 |
| GO:0042551 | neuron maturation | 2.00e-02 | 1.00e+00 | 5.628 | 1 | 1 | 11 |
| GO:0045651 | positive regulation of macrophage differentiation | 2.00e-02 | 1.00e+00 | 5.628 | 1 | 1 | 11 |
| GO:0045120 | pronucleus | 2.00e-02 | 1.00e+00 | 5.628 | 1 | 1 | 11 |
| GO:0035518 | histone H2A monoubiquitination | 2.00e-02 | 1.00e+00 | 5.628 | 1 | 2 | 11 |
| GO:0002011 | morphogenesis of an epithelial sheet | 2.00e-02 | 1.00e+00 | 5.628 | 1 | 1 | 11 |
| GO:0021819 | layer formation in cerebral cortex | 2.00e-02 | 1.00e+00 | 5.628 | 1 | 1 | 11 |
| GO:0051056 | regulation of small GTPase mediated signal transduction | 2.14e-02 | 1.00e+00 | 3.145 | 2 | 3 | 123 |
| GO:0007051 | spindle organization | 2.18e-02 | 1.00e+00 | 5.503 | 1 | 1 | 12 |
| GO:0051146 | striated muscle cell differentiation | 2.18e-02 | 1.00e+00 | 5.503 | 1 | 2 | 12 |
| GO:0043149 | stress fiber assembly | 2.18e-02 | 1.00e+00 | 5.503 | 1 | 2 | 12 |
| GO:0030140 | trans-Golgi network transport vesicle | 2.18e-02 | 1.00e+00 | 5.503 | 1 | 1 | 12 |
| GO:0072583 | clathrin-mediated endocytosis | 2.18e-02 | 1.00e+00 | 5.503 | 1 | 1 | 12 |
| GO:0047496 | vesicle transport along microtubule | 2.18e-02 | 1.00e+00 | 5.503 | 1 | 1 | 12 |
| GO:0055003 | cardiac myofibril assembly | 2.18e-02 | 1.00e+00 | 5.503 | 1 | 1 | 12 |
| GO:0042789 | mRNA transcription from RNA polymerase II promoter | 2.36e-02 | 1.00e+00 | 5.387 | 1 | 1 | 13 |
| GO:0001671 | ATPase activator activity | 2.36e-02 | 1.00e+00 | 5.387 | 1 | 1 | 13 |
| GO:0043488 | regulation of mRNA stability | 2.36e-02 | 1.00e+00 | 5.387 | 1 | 1 | 13 |
| GO:0005662 | DNA replication factor A complex | 2.36e-02 | 1.00e+00 | 5.387 | 1 | 1 | 13 |
| GO:1900087 | positive regulation of G1/S transition of mitotic cell cycle | 2.36e-02 | 1.00e+00 | 5.387 | 1 | 1 | 13 |
| GO:0060314 | regulation of ryanodine-sensitive calcium-release channel activity | 2.36e-02 | 1.00e+00 | 5.387 | 1 | 1 | 13 |
| GO:0035855 | megakaryocyte development | 2.36e-02 | 1.00e+00 | 5.387 | 1 | 1 | 13 |
| GO:0005813 | centrosome | 2.38e-02 | 1.00e+00 | 2.267 | 3 | 9 | 339 |
| GO:0000790 | nuclear chromatin | 2.47e-02 | 1.00e+00 | 3.032 | 2 | 2 | 133 |
| GO:0035371 | microtubule plus-end | 2.54e-02 | 1.00e+00 | 5.280 | 1 | 1 | 14 |
| GO:0031333 | negative regulation of protein complex assembly | 2.54e-02 | 1.00e+00 | 5.280 | 1 | 1 | 14 |
| GO:0060292 | long term synaptic depression | 2.54e-02 | 1.00e+00 | 5.280 | 1 | 1 | 14 |
| GO:0048488 | synaptic vesicle endocytosis | 2.54e-02 | 1.00e+00 | 5.280 | 1 | 1 | 14 |
| GO:0035267 | NuA4 histone acetyltransferase complex | 2.54e-02 | 1.00e+00 | 5.280 | 1 | 2 | 14 |
| GO:0031996 | thioesterase binding | 2.54e-02 | 1.00e+00 | 5.280 | 1 | 1 | 14 |
| GO:0034185 | apolipoprotein binding | 2.54e-02 | 1.00e+00 | 5.280 | 1 | 1 | 14 |
| GO:0031334 | positive regulation of protein complex assembly | 2.54e-02 | 1.00e+00 | 5.280 | 1 | 1 | 14 |
| GO:0000086 | G2/M transition of mitotic cell cycle | 2.61e-02 | 1.00e+00 | 2.990 | 2 | 6 | 137 |
| GO:0030131 | clathrin adaptor complex | 2.72e-02 | 1.00e+00 | 5.181 | 1 | 1 | 15 |
| GO:0002042 | cell migration involved in sprouting angiogenesis | 2.72e-02 | 1.00e+00 | 5.181 | 1 | 1 | 15 |
| GO:0030225 | macrophage differentiation | 2.72e-02 | 1.00e+00 | 5.181 | 1 | 1 | 15 |
| GO:0016514 | SWI/SNF complex | 2.72e-02 | 1.00e+00 | 5.181 | 1 | 1 | 15 |
| GO:0045987 | positive regulation of smooth muscle contraction | 2.72e-02 | 1.00e+00 | 5.181 | 1 | 1 | 15 |
| GO:0045445 | myoblast differentiation | 2.72e-02 | 1.00e+00 | 5.181 | 1 | 1 | 15 |
| GO:0048821 | erythrocyte development | 2.72e-02 | 1.00e+00 | 5.181 | 1 | 2 | 15 |
| GO:0016010 | dystrophin-associated glycoprotein complex | 2.72e-02 | 1.00e+00 | 5.181 | 1 | 1 | 15 |
| GO:2000114 | regulation of establishment of cell polarity | 2.72e-02 | 1.00e+00 | 5.181 | 1 | 1 | 15 |
| GO:0051233 | spindle midzone | 2.72e-02 | 1.00e+00 | 5.181 | 1 | 2 | 15 |
| GO:0048854 | brain morphogenesis | 2.72e-02 | 1.00e+00 | 5.181 | 1 | 1 | 15 |
| GO:0060347 | heart trabecula formation | 2.72e-02 | 1.00e+00 | 5.181 | 1 | 1 | 15 |
| GO:0032839 | dendrite cytoplasm | 2.90e-02 | 1.00e+00 | 5.088 | 1 | 1 | 16 |
| GO:0007405 | neuroblast proliferation | 2.90e-02 | 1.00e+00 | 5.088 | 1 | 1 | 16 |
| GO:0032012 | regulation of ARF protein signal transduction | 2.90e-02 | 1.00e+00 | 5.088 | 1 | 1 | 16 |
| GO:0019226 | transmission of nerve impulse | 2.90e-02 | 1.00e+00 | 5.088 | 1 | 1 | 16 |
| GO:0000132 | establishment of mitotic spindle orientation | 2.90e-02 | 1.00e+00 | 5.088 | 1 | 2 | 16 |
| GO:0042176 | regulation of protein catabolic process | 2.90e-02 | 1.00e+00 | 5.088 | 1 | 4 | 16 |
| GO:0033137 | negative regulation of peptidyl-serine phosphorylation | 2.90e-02 | 1.00e+00 | 5.088 | 1 | 1 | 16 |
| GO:0030220 | platelet formation | 2.90e-02 | 1.00e+00 | 5.088 | 1 | 1 | 16 |
| GO:0043623 | cellular protein complex assembly | 2.90e-02 | 1.00e+00 | 5.088 | 1 | 1 | 16 |
| GO:0046716 | muscle cell cellular homeostasis | 2.90e-02 | 1.00e+00 | 5.088 | 1 | 1 | 16 |
| GO:0050775 | positive regulation of dendrite morphogenesis | 2.90e-02 | 1.00e+00 | 5.088 | 1 | 1 | 16 |
| GO:0008285 | negative regulation of cell proliferation | 2.92e-02 | 1.00e+00 | 2.153 | 3 | 3 | 367 |
| GO:0005794 | Golgi apparatus | 3.00e-02 | 1.00e+00 | 1.743 | 4 | 8 | 650 |
| GO:0006457 | protein folding | 3.05e-02 | 1.00e+00 | 2.868 | 2 | 3 | 149 |
| GO:0035255 | ionotropic glutamate receptor binding | 3.08e-02 | 1.00e+00 | 5.000 | 1 | 1 | 17 |
| GO:0010880 | regulation of release of sequestered calcium ion into cytosol by sarcoplasmic reticulum | 3.08e-02 | 1.00e+00 | 5.000 | 1 | 1 | 17 |
| GO:0043274 | phospholipase binding | 3.08e-02 | 1.00e+00 | 5.000 | 1 | 1 | 17 |
| GO:0008306 | associative learning | 3.08e-02 | 1.00e+00 | 5.000 | 1 | 1 | 17 |
| GO:0031527 | filopodium membrane | 3.08e-02 | 1.00e+00 | 5.000 | 1 | 1 | 17 |
| GO:0001829 | trophectodermal cell differentiation | 3.08e-02 | 1.00e+00 | 5.000 | 1 | 1 | 17 |
| GO:0015935 | small ribosomal subunit | 3.08e-02 | 1.00e+00 | 5.000 | 1 | 9 | 17 |
| GO:0030742 | GTP-dependent protein binding | 3.08e-02 | 1.00e+00 | 5.000 | 1 | 1 | 17 |
| GO:0070062 | extracellular vesicular exosome | 3.24e-02 | 1.00e+00 | 0.961 | 9 | 51 | 2516 |
| GO:0006612 | protein targeting to membrane | 3.26e-02 | 1.00e+00 | 4.918 | 1 | 2 | 18 |
| GO:0045773 | positive regulation of axon extension | 3.26e-02 | 1.00e+00 | 4.918 | 1 | 1 | 18 |
| GO:0005086 | ARF guanyl-nucleotide exchange factor activity | 3.26e-02 | 1.00e+00 | 4.918 | 1 | 1 | 18 |
| GO:0090316 | positive regulation of intracellular protein transport | 3.26e-02 | 1.00e+00 | 4.918 | 1 | 1 | 18 |
| GO:0030532 | small nuclear ribonucleoprotein complex | 3.26e-02 | 1.00e+00 | 4.918 | 1 | 1 | 18 |
| GO:0005788 | endoplasmic reticulum lumen | 3.35e-02 | 1.00e+00 | 2.793 | 2 | 2 | 157 |
| GO:0007088 | regulation of mitosis | 3.44e-02 | 1.00e+00 | 4.840 | 1 | 1 | 19 |
| GO:0010881 | regulation of cardiac muscle contraction by regulation of the release of sequestered calcium ion | 3.44e-02 | 1.00e+00 | 4.840 | 1 | 1 | 19 |
| GO:0030866 | cortical actin cytoskeleton organization | 3.44e-02 | 1.00e+00 | 4.840 | 1 | 1 | 19 |
| GO:0045121 | membrane raft | 3.51e-02 | 1.00e+00 | 2.757 | 2 | 3 | 161 |
| GO:0000278 | mitotic cell cycle | 3.59e-02 | 1.00e+00 | 2.036 | 3 | 16 | 398 |
| GO:2000134 | negative regulation of G1/S transition of mitotic cell cycle | 3.61e-02 | 1.00e+00 | 4.766 | 1 | 1 | 20 |
| GO:0090398 | cellular senescence | 3.61e-02 | 1.00e+00 | 4.766 | 1 | 1 | 20 |
| GO:0030544 | Hsp70 protein binding | 3.61e-02 | 1.00e+00 | 4.766 | 1 | 1 | 20 |
| GO:0043473 | pigmentation | 3.61e-02 | 1.00e+00 | 4.766 | 1 | 2 | 20 |
| GO:0015035 | protein disulfide oxidoreductase activity | 3.79e-02 | 1.00e+00 | 4.695 | 1 | 1 | 21 |
| GO:0043034 | costamere | 3.79e-02 | 1.00e+00 | 4.695 | 1 | 1 | 21 |
| GO:0046847 | filopodium assembly | 3.79e-02 | 1.00e+00 | 4.695 | 1 | 1 | 21 |
| GO:0005790 | smooth endoplasmic reticulum | 3.97e-02 | 1.00e+00 | 4.628 | 1 | 1 | 22 |
| GO:0031435 | mitogen-activated protein kinase kinase kinase binding | 3.97e-02 | 1.00e+00 | 4.628 | 1 | 1 | 22 |
| GO:0000083 | regulation of transcription involved in G1/S transition of mitotic cell cycle | 3.97e-02 | 1.00e+00 | 4.628 | 1 | 1 | 22 |
| GO:0030863 | cortical cytoskeleton | 3.97e-02 | 1.00e+00 | 4.628 | 1 | 1 | 22 |
| GO:1900026 | positive regulation of substrate adhesion-dependent cell spreading | 4.15e-02 | 1.00e+00 | 4.564 | 1 | 1 | 23 |
| GO:0002040 | sprouting angiogenesis | 4.15e-02 | 1.00e+00 | 4.564 | 1 | 1 | 23 |
| GO:0045879 | negative regulation of smoothened signaling pathway | 4.15e-02 | 1.00e+00 | 4.564 | 1 | 1 | 23 |
| GO:0051017 | actin filament bundle assembly | 4.15e-02 | 1.00e+00 | 4.564 | 1 | 1 | 23 |
| GO:0008060 | ARF GTPase activator activity | 4.15e-02 | 1.00e+00 | 4.564 | 1 | 1 | 23 |
| GO:0001892 | embryonic placenta development | 4.15e-02 | 1.00e+00 | 4.564 | 1 | 1 | 23 |
| GO:0051297 | centrosome organization | 4.15e-02 | 1.00e+00 | 4.564 | 1 | 2 | 23 |
| GO:0043044 | ATP-dependent chromatin remodeling | 4.15e-02 | 1.00e+00 | 4.564 | 1 | 1 | 23 |
| GO:0045211 | postsynaptic membrane | 4.17e-02 | 1.00e+00 | 2.620 | 2 | 3 | 177 |
| GO:0031625 | ubiquitin protein ligase binding | 4.30e-02 | 1.00e+00 | 2.596 | 2 | 4 | 180 |
| GO:0008135 | translation factor activity, nucleic acid binding | 4.32e-02 | 1.00e+00 | 4.503 | 1 | 4 | 24 |
| GO:0005761 | mitochondrial ribosome | 4.32e-02 | 1.00e+00 | 4.503 | 1 | 1 | 24 |
| GO:0070059 | intrinsic apoptotic signaling pathway in response to endoplasmic reticulum stress | 4.32e-02 | 1.00e+00 | 4.503 | 1 | 1 | 24 |
| GO:0007163 | establishment or maintenance of cell polarity | 4.32e-02 | 1.00e+00 | 4.503 | 1 | 2 | 24 |
| GO:0046329 | negative regulation of JNK cascade | 4.32e-02 | 1.00e+00 | 4.503 | 1 | 1 | 24 |
| GO:0005100 | Rho GTPase activator activity | 4.32e-02 | 1.00e+00 | 4.503 | 1 | 1 | 24 |
| GO:0032781 | positive regulation of ATPase activity | 4.50e-02 | 1.00e+00 | 4.444 | 1 | 1 | 25 |
| GO:0046326 | positive regulation of glucose import | 4.50e-02 | 1.00e+00 | 4.444 | 1 | 1 | 25 |
| GO:0045931 | positive regulation of mitotic cell cycle | 4.67e-02 | 1.00e+00 | 4.387 | 1 | 2 | 26 |
| GO:0045214 | sarcomere organization | 4.67e-02 | 1.00e+00 | 4.387 | 1 | 1 | 26 |
| GO:0045859 | regulation of protein kinase activity | 4.67e-02 | 1.00e+00 | 4.387 | 1 | 1 | 26 |
| GO:0048589 | developmental growth | 4.67e-02 | 1.00e+00 | 4.387 | 1 | 1 | 26 |
| GO:0001707 | mesoderm formation | 4.67e-02 | 1.00e+00 | 4.387 | 1 | 1 | 26 |
| GO:0043552 | positive regulation of phosphatidylinositol 3-kinase activity | 4.67e-02 | 1.00e+00 | 4.387 | 1 | 2 | 26 |
| GO:0071556 | integral component of lumenal side of endoplasmic reticulum membrane | 4.67e-02 | 1.00e+00 | 4.387 | 1 | 1 | 26 |
| GO:0043547 | positive regulation of GTPase activity | 4.74e-02 | 1.00e+00 | 2.518 | 2 | 4 | 190 |
| GO:0051149 | positive regulation of muscle cell differentiation | 4.85e-02 | 1.00e+00 | 4.333 | 1 | 2 | 27 |
| GO:0032720 | negative regulation of tumor necrosis factor production | 4.85e-02 | 1.00e+00 | 4.333 | 1 | 1 | 27 |
| GO:0007616 | long-term memory | 4.85e-02 | 1.00e+00 | 4.333 | 1 | 1 | 27 |
| GO:0005083 | small GTPase regulator activity | 4.85e-02 | 1.00e+00 | 4.333 | 1 | 1 | 27 |
| GO:0031424 | keratinization | 4.85e-02 | 1.00e+00 | 4.333 | 1 | 1 | 27 |
| GO:0048565 | digestive tract development | 4.85e-02 | 1.00e+00 | 4.333 | 1 | 1 | 27 |
| GO:0031069 | hair follicle morphogenesis | 4.85e-02 | 1.00e+00 | 4.333 | 1 | 1 | 27 |
| GO:0032467 | positive regulation of cytokinesis | 5.03e-02 | 1.00e+00 | 4.280 | 1 | 1 | 28 |
| GO:0031492 | nucleosomal DNA binding | 5.03e-02 | 1.00e+00 | 4.280 | 1 | 1 | 28 |
| GO:0019894 | kinesin binding | 5.03e-02 | 1.00e+00 | 4.280 | 1 | 1 | 28 |
| GO:0005875 | microtubule associated complex | 5.03e-02 | 1.00e+00 | 4.280 | 1 | 1 | 28 |
| GO:0045597 | positive regulation of cell differentiation | 5.03e-02 | 1.00e+00 | 4.280 | 1 | 1 | 28 |
| GO:0007017 | microtubule-based process | 5.03e-02 | 1.00e+00 | 4.280 | 1 | 2 | 28 |
| GO:0031252 | cell leading edge | 5.20e-02 | 1.00e+00 | 4.230 | 1 | 3 | 29 |
| GO:0003730 | mRNA 3'-UTR binding | 5.20e-02 | 1.00e+00 | 4.230 | 1 | 1 | 29 |
| GO:0034605 | cellular response to heat | 5.20e-02 | 1.00e+00 | 4.230 | 1 | 1 | 29 |
| GO:0072686 | mitotic spindle | 5.20e-02 | 1.00e+00 | 4.230 | 1 | 1 | 29 |
| GO:0003924 | GTPase activity | 5.33e-02 | 1.00e+00 | 2.422 | 2 | 6 | 203 |
| GO:0010977 | negative regulation of neuron projection development | 5.37e-02 | 1.00e+00 | 4.181 | 1 | 2 | 30 |
| GO:0007346 | regulation of mitotic cell cycle | 5.37e-02 | 1.00e+00 | 4.181 | 1 | 3 | 30 |
| GO:0031647 | regulation of protein stability | 5.37e-02 | 1.00e+00 | 4.181 | 1 | 1 | 30 |
| GO:0032956 | regulation of actin cytoskeleton organization | 5.37e-02 | 1.00e+00 | 4.181 | 1 | 1 | 30 |
| GO:0002027 | regulation of heart rate | 5.37e-02 | 1.00e+00 | 4.181 | 1 | 2 | 30 |
| GO:0040018 | positive regulation of multicellular organism growth | 5.37e-02 | 1.00e+00 | 4.181 | 1 | 1 | 30 |
| GO:0010494 | cytoplasmic stress granule | 5.37e-02 | 1.00e+00 | 4.181 | 1 | 2 | 30 |
| GO:0061077 | chaperone-mediated protein folding | 5.55e-02 | 1.00e+00 | 4.133 | 1 | 1 | 31 |
| GO:0007093 | mitotic cell cycle checkpoint | 5.55e-02 | 1.00e+00 | 4.133 | 1 | 2 | 31 |
| GO:0070830 | tight junction assembly | 5.90e-02 | 1.00e+00 | 4.043 | 1 | 1 | 33 |
| GO:0045944 | positive regulation of transcription from RNA polymerase II promoter | 5.95e-02 | 1.00e+00 | 1.424 | 4 | 11 | 811 |
| GO:0001569 | patterning of blood vessels | 6.07e-02 | 1.00e+00 | 4.000 | 1 | 1 | 34 |
| GO:0007611 | learning or memory | 6.07e-02 | 1.00e+00 | 4.000 | 1 | 2 | 34 |
| GO:0042692 | muscle cell differentiation | 6.07e-02 | 1.00e+00 | 4.000 | 1 | 2 | 34 |
| GO:0044267 | cellular protein metabolic process | 6.15e-02 | 1.00e+00 | 1.721 | 3 | 29 | 495 |
| GO:0006184 | GTP catabolic process | 6.15e-02 | 1.00e+00 | 2.306 | 2 | 6 | 220 |
| GO:0048666 | neuron development | 6.24e-02 | 1.00e+00 | 3.958 | 1 | 1 | 35 |
| GO:0071333 | cellular response to glucose stimulus | 6.24e-02 | 1.00e+00 | 3.958 | 1 | 2 | 35 |
| GO:0009725 | response to hormone | 6.24e-02 | 1.00e+00 | 3.958 | 1 | 1 | 35 |
| GO:0005886 | plasma membrane | 6.30e-02 | 1.00e+00 | 0.789 | 9 | 24 | 2834 |
| GO:0051402 | neuron apoptotic process | 6.42e-02 | 1.00e+00 | 3.918 | 1 | 1 | 36 |
| GO:0030838 | positive regulation of actin filament polymerization | 6.42e-02 | 1.00e+00 | 3.918 | 1 | 1 | 36 |
| GO:0030433 | ER-associated ubiquitin-dependent protein catabolic process | 6.42e-02 | 1.00e+00 | 3.918 | 1 | 1 | 36 |
| GO:0001895 | retina homeostasis | 6.42e-02 | 1.00e+00 | 3.918 | 1 | 1 | 36 |
| GO:0042059 | negative regulation of epidermal growth factor receptor signaling pathway | 6.42e-02 | 1.00e+00 | 3.918 | 1 | 3 | 36 |
| GO:0001102 | RNA polymerase II activating transcription factor binding | 6.59e-02 | 1.00e+00 | 3.878 | 1 | 1 | 37 |
| GO:0051084 | 'de novo' posttranslational protein folding | 6.59e-02 | 1.00e+00 | 3.878 | 1 | 2 | 37 |
| GO:0005791 | rough endoplasmic reticulum | 6.59e-02 | 1.00e+00 | 3.878 | 1 | 1 | 37 |
| GO:0007067 | mitotic nuclear division | 6.70e-02 | 1.00e+00 | 2.236 | 2 | 7 | 231 |
| GO:0001568 | blood vessel development | 6.76e-02 | 1.00e+00 | 3.840 | 1 | 2 | 38 |
| GO:0070527 | platelet aggregation | 6.76e-02 | 1.00e+00 | 3.840 | 1 | 1 | 38 |
| GO:0050681 | androgen receptor binding | 6.76e-02 | 1.00e+00 | 3.840 | 1 | 2 | 38 |
| GO:0030049 | muscle filament sliding | 6.76e-02 | 1.00e+00 | 3.840 | 1 | 2 | 38 |
| GO:0045740 | positive regulation of DNA replication | 6.76e-02 | 1.00e+00 | 3.840 | 1 | 1 | 38 |
| GO:0006412 | translation | 6.90e-02 | 1.00e+00 | 2.211 | 2 | 29 | 235 |
| GO:0031490 | chromatin DNA binding | 6.93e-02 | 1.00e+00 | 3.802 | 1 | 1 | 39 |
| GO:0060048 | cardiac muscle contraction | 6.93e-02 | 1.00e+00 | 3.802 | 1 | 1 | 39 |
| GO:0001933 | negative regulation of protein phosphorylation | 6.93e-02 | 1.00e+00 | 3.802 | 1 | 1 | 39 |
| GO:0048471 | perinuclear region of cytoplasm | 7.01e-02 | 1.00e+00 | 1.642 | 3 | 9 | 523 |
| GO:0001228 | RNA polymerase II transcription regulatory region sequence-specific DNA binding transcription factor activity involved in positive regulation of transcription | 7.10e-02 | 1.00e+00 | 3.766 | 1 | 1 | 40 |
| GO:0007015 | actin filament organization | 7.10e-02 | 1.00e+00 | 3.766 | 1 | 1 | 40 |
| GO:0030425 | dendrite | 7.16e-02 | 1.00e+00 | 2.181 | 2 | 3 | 240 |
| GO:0051117 | ATPase binding | 7.27e-02 | 1.00e+00 | 3.730 | 1 | 2 | 41 |
| GO:0008307 | structural constituent of muscle | 7.27e-02 | 1.00e+00 | 3.730 | 1 | 1 | 41 |
| GO:0017148 | negative regulation of translation | 7.27e-02 | 1.00e+00 | 3.730 | 1 | 1 | 41 |
| GO:0030521 | androgen receptor signaling pathway | 7.27e-02 | 1.00e+00 | 3.730 | 1 | 3 | 41 |
| GO:0050885 | neuromuscular process controlling balance | 7.27e-02 | 1.00e+00 | 3.730 | 1 | 1 | 41 |
| GO:0021987 | cerebral cortex development | 7.45e-02 | 1.00e+00 | 3.695 | 1 | 1 | 42 |
| GO:0030155 | regulation of cell adhesion | 7.45e-02 | 1.00e+00 | 3.695 | 1 | 1 | 42 |
| GO:0035914 | skeletal muscle cell differentiation | 7.45e-02 | 1.00e+00 | 3.695 | 1 | 1 | 42 |
| GO:0005871 | kinesin complex | 7.79e-02 | 1.00e+00 | 3.628 | 1 | 1 | 44 |
| GO:0050896 | response to stimulus | 7.79e-02 | 1.00e+00 | 3.628 | 1 | 1 | 44 |
| GO:0016328 | lateral plasma membrane | 8.13e-02 | 1.00e+00 | 3.564 | 1 | 3 | 46 |
| GO:0043525 | positive regulation of neuron apoptotic process | 8.13e-02 | 1.00e+00 | 3.564 | 1 | 2 | 46 |
| GO:0001047 | core promoter binding | 8.13e-02 | 1.00e+00 | 3.564 | 1 | 1 | 46 |
| GO:0005884 | actin filament | 8.13e-02 | 1.00e+00 | 3.564 | 1 | 2 | 46 |
| GO:0001947 | heart looping | 8.63e-02 | 1.00e+00 | 3.473 | 1 | 1 | 49 |
| GO:0003743 | translation initiation factor activity | 8.63e-02 | 1.00e+00 | 3.473 | 1 | 5 | 49 |
| GO:0034097 | response to cytokine | 8.80e-02 | 1.00e+00 | 3.444 | 1 | 1 | 50 |
| GO:0001948 | glycoprotein binding | 8.80e-02 | 1.00e+00 | 3.444 | 1 | 2 | 50 |
| GO:0003684 | damaged DNA binding | 8.97e-02 | 1.00e+00 | 3.415 | 1 | 2 | 51 |
| GO:0045732 | positive regulation of protein catabolic process | 8.97e-02 | 1.00e+00 | 3.415 | 1 | 1 | 51 |
| GO:0045454 | cell redox homeostasis | 9.14e-02 | 1.00e+00 | 3.387 | 1 | 1 | 52 |
| GO:0034976 | response to endoplasmic reticulum stress | 9.14e-02 | 1.00e+00 | 3.387 | 1 | 1 | 52 |
| GO:0016042 | lipid catabolic process | 9.14e-02 | 1.00e+00 | 3.387 | 1 | 2 | 52 |
| GO:0006952 | defense response | 9.30e-02 | 1.00e+00 | 3.360 | 1 | 1 | 53 |
| GO:0019900 | kinase binding | 9.47e-02 | 1.00e+00 | 3.333 | 1 | 3 | 54 |
| GO:0000980 | RNA polymerase II distal enhancer sequence-specific DNA binding | 9.47e-02 | 1.00e+00 | 3.333 | 1 | 1 | 54 |
| GO:0050680 | negative regulation of epithelial cell proliferation | 9.47e-02 | 1.00e+00 | 3.333 | 1 | 1 | 54 |
| GO:0000226 | microtubule cytoskeleton organization | 9.64e-02 | 1.00e+00 | 3.306 | 1 | 1 | 55 |
| GO:0046330 | positive regulation of JNK cascade | 9.64e-02 | 1.00e+00 | 3.306 | 1 | 1 | 55 |
| GO:0006888 | ER to Golgi vesicle-mediated transport | 9.64e-02 | 1.00e+00 | 3.306 | 1 | 1 | 55 |
| GO:0000932 | cytoplasmic mRNA processing body | 9.81e-02 | 1.00e+00 | 3.280 | 1 | 1 | 56 |
| GO:0007264 | small GTPase mediated signal transduction | 9.89e-02 | 1.00e+00 | 1.908 | 2 | 7 | 290 |
| GO:0005793 | endoplasmic reticulum-Golgi intermediate compartment | 9.97e-02 | 1.00e+00 | 3.255 | 1 | 1 | 57 |
| GO:0012505 | endomembrane system | 9.97e-02 | 1.00e+00 | 3.255 | 1 | 1 | 57 |
| GO:0005783 | endoplasmic reticulum | 1.00e-01 | 1.00e+00 | 1.420 | 3 | 6 | 610 |
| GO:0051087 | chaperone binding | 1.03e-01 | 1.00e+00 | 3.205 | 1 | 1 | 59 |
| GO:0045216 | cell-cell junction organization | 1.03e-01 | 1.00e+00 | 3.205 | 1 | 2 | 59 |
| GO:0005840 | ribosome | 1.03e-01 | 1.00e+00 | 3.205 | 1 | 10 | 59 |
| GO:0043433 | negative regulation of sequence-specific DNA binding transcription factor activity | 1.03e-01 | 1.00e+00 | 3.205 | 1 | 1 | 59 |
| GO:0010976 | positive regulation of neuron projection development | 1.05e-01 | 1.00e+00 | 3.181 | 1 | 2 | 60 |
| GO:0003705 | RNA polymerase II distal enhancer sequence-specific DNA binding transcription factor activity | 1.06e-01 | 1.00e+00 | 3.157 | 1 | 1 | 61 |
| GO:0033138 | positive regulation of peptidyl-serine phosphorylation | 1.06e-01 | 1.00e+00 | 3.157 | 1 | 1 | 61 |
| GO:0001078 | RNA polymerase II core promoter proximal region sequence-specific DNA binding transcription factor activity involved in negative regulation of transcription | 1.08e-01 | 1.00e+00 | 3.133 | 1 | 1 | 62 |
| GO:0000776 | kinetochore | 1.10e-01 | 1.00e+00 | 3.110 | 1 | 3 | 63 |
| GO:0032869 | cellular response to insulin stimulus | 1.11e-01 | 1.00e+00 | 3.088 | 1 | 2 | 64 |
| GO:0005856 | cytoskeleton | 1.11e-01 | 1.00e+00 | 1.807 | 2 | 6 | 311 |
| GO:0006469 | negative regulation of protein kinase activity | 1.13e-01 | 1.00e+00 | 3.065 | 1 | 1 | 65 |
| GO:0071260 | cellular response to mechanical stimulus | 1.15e-01 | 1.00e+00 | 3.043 | 1 | 3 | 66 |
| GO:0009636 | response to toxic substance | 1.15e-01 | 1.00e+00 | 3.043 | 1 | 1 | 66 |
| GO:0031295 | T cell costimulation | 1.16e-01 | 1.00e+00 | 3.021 | 1 | 3 | 67 |
| GO:0030141 | secretory granule | 1.16e-01 | 1.00e+00 | 3.021 | 1 | 2 | 67 |
| GO:0019901 | protein kinase binding | 1.17e-01 | 1.00e+00 | 1.766 | 2 | 9 | 320 |
| GO:0006338 | chromatin remodeling | 1.18e-01 | 1.00e+00 | 3.000 | 1 | 2 | 68 |
| GO:0045666 | positive regulation of neuron differentiation | 1.18e-01 | 1.00e+00 | 3.000 | 1 | 1 | 68 |
| GO:0050790 | regulation of catalytic activity | 1.19e-01 | 1.00e+00 | 2.979 | 1 | 1 | 69 |
| GO:0006887 | exocytosis | 1.19e-01 | 1.00e+00 | 2.979 | 1 | 1 | 69 |
| GO:0007411 | axon guidance | 1.21e-01 | 1.00e+00 | 1.734 | 2 | 3 | 327 |
| GO:0035264 | multicellular organism growth | 1.21e-01 | 1.00e+00 | 2.958 | 1 | 1 | 70 |
| GO:0005525 | GTP binding | 1.21e-01 | 1.00e+00 | 1.730 | 2 | 6 | 328 |
| GO:0034329 | cell junction assembly | 1.23e-01 | 1.00e+00 | 2.938 | 1 | 2 | 71 |
| GO:0042383 | sarcolemma | 1.23e-01 | 1.00e+00 | 2.938 | 1 | 2 | 71 |
| GO:0043231 | intracellular membrane-bounded organelle | 1.24e-01 | 1.00e+00 | 1.713 | 2 | 3 | 332 |
| GO:0003682 | chromatin binding | 1.25e-01 | 1.00e+00 | 1.704 | 2 | 4 | 334 |
| GO:0000785 | chromatin | 1.26e-01 | 1.00e+00 | 2.898 | 1 | 2 | 73 |
| GO:0005085 | guanyl-nucleotide exchange factor activity | 1.28e-01 | 1.00e+00 | 2.878 | 1 | 1 | 74 |
| GO:0007265 | Ras protein signal transduction | 1.29e-01 | 1.00e+00 | 2.859 | 1 | 3 | 75 |
| GO:0060070 | canonical Wnt signaling pathway | 1.29e-01 | 1.00e+00 | 2.859 | 1 | 2 | 75 |
| GO:0044325 | ion channel binding | 1.32e-01 | 1.00e+00 | 2.821 | 1 | 3 | 77 |
| GO:0008584 | male gonad development | 1.32e-01 | 1.00e+00 | 2.821 | 1 | 1 | 77 |
| GO:0017137 | Rab GTPase binding | 1.34e-01 | 1.00e+00 | 2.802 | 1 | 1 | 78 |
| GO:0071013 | catalytic step 2 spliceosome | 1.36e-01 | 1.00e+00 | 2.784 | 1 | 1 | 79 |
| GO:0010629 | negative regulation of gene expression | 1.37e-01 | 1.00e+00 | 2.766 | 1 | 2 | 80 |
| GO:0003723 | RNA binding | 1.38e-01 | 1.00e+00 | 1.616 | 2 | 10 | 355 |
| GO:0015031 | protein transport | 1.39e-01 | 1.00e+00 | 1.608 | 2 | 4 | 357 |
| GO:0030054 | cell junction | 1.39e-01 | 1.00e+00 | 1.612 | 2 | 4 | 356 |
| GO:0045177 | apical part of cell | 1.40e-01 | 1.00e+00 | 2.730 | 1 | 1 | 82 |
| GO:0030336 | negative regulation of cell migration | 1.42e-01 | 1.00e+00 | 2.713 | 1 | 2 | 83 |
| GO:0007517 | muscle organ development | 1.42e-01 | 1.00e+00 | 2.713 | 1 | 1 | 83 |
| GO:0043197 | dendritic spine | 1.42e-01 | 1.00e+00 | 2.713 | 1 | 2 | 83 |
| GO:0005179 | hormone activity | 1.44e-01 | 1.00e+00 | 2.695 | 1 | 1 | 84 |
| GO:0047485 | protein N-terminus binding | 1.47e-01 | 1.00e+00 | 2.661 | 1 | 1 | 86 |
| GO:0005925 | focal adhesion | 1.48e-01 | 1.00e+00 | 1.556 | 2 | 23 | 370 |
| GO:0007160 | cell-matrix adhesion | 1.50e-01 | 1.00e+00 | 2.628 | 1 | 1 | 88 |
| GO:0032321 | positive regulation of Rho GTPase activity | 1.50e-01 | 1.00e+00 | 2.628 | 1 | 1 | 88 |
| GO:0000922 | spindle pole | 1.55e-01 | 1.00e+00 | 2.580 | 1 | 5 | 91 |
| GO:0018279 | protein N-linked glycosylation via asparagine | 1.55e-01 | 1.00e+00 | 2.580 | 1 | 1 | 91 |
| GO:0006928 | cellular component movement | 1.56e-01 | 1.00e+00 | 2.564 | 1 | 4 | 92 |
| GO:0016363 | nuclear matrix | 1.56e-01 | 1.00e+00 | 2.564 | 1 | 4 | 92 |
| GO:0016605 | PML body | 1.56e-01 | 1.00e+00 | 2.564 | 1 | 2 | 92 |
| GO:0042470 | melanosome | 1.56e-01 | 1.00e+00 | 2.564 | 1 | 2 | 92 |
| GO:0003700 | sequence-specific DNA binding transcription factor activity | 1.57e-01 | 1.00e+00 | 1.126 | 3 | 9 | 748 |
| GO:0051091 | positive regulation of sequence-specific DNA binding transcription factor activity | 1.58e-01 | 1.00e+00 | 2.548 | 1 | 1 | 93 |
| GO:0002474 | antigen processing and presentation of peptide antigen via MHC class I | 1.59e-01 | 1.00e+00 | 2.533 | 1 | 7 | 94 |
| GO:0051082 | unfolded protein binding | 1.61e-01 | 1.00e+00 | 2.518 | 1 | 1 | 95 |
| GO:0030426 | growth cone | 1.64e-01 | 1.00e+00 | 2.488 | 1 | 3 | 97 |
| GO:0004888 | transmembrane signaling receptor activity | 1.73e-01 | 1.00e+00 | 2.401 | 1 | 1 | 103 |
| GO:0005096 | GTPase activator activity | 1.81e-01 | 1.00e+00 | 2.333 | 1 | 3 | 108 |
| GO:0030496 | midbody | 1.82e-01 | 1.00e+00 | 2.319 | 1 | 4 | 109 |
| GO:0005938 | cell cortex | 1.82e-01 | 1.00e+00 | 2.319 | 1 | 1 | 109 |
| GO:0005815 | microtubule organizing center | 1.84e-01 | 1.00e+00 | 2.306 | 1 | 4 | 110 |
| GO:0030529 | ribonucleoprotein complex | 1.90e-01 | 1.00e+00 | 2.255 | 1 | 8 | 114 |
| GO:0005819 | spindle | 1.90e-01 | 1.00e+00 | 2.255 | 1 | 4 | 114 |
| GO:0072562 | blood microparticle | 1.93e-01 | 1.00e+00 | 2.230 | 1 | 1 | 116 |
| GO:0005802 | trans-Golgi network | 1.93e-01 | 1.00e+00 | 2.230 | 1 | 2 | 116 |
| GO:0005635 | nuclear envelope | 1.93e-01 | 1.00e+00 | 2.230 | 1 | 2 | 116 |
| GO:0032496 | response to lipopolysaccharide | 2.02e-01 | 1.00e+00 | 2.157 | 1 | 1 | 122 |
| GO:0016192 | vesicle-mediated transport | 2.02e-01 | 1.00e+00 | 2.157 | 1 | 1 | 122 |
| GO:0006325 | chromatin organization | 2.03e-01 | 1.00e+00 | 2.145 | 1 | 2 | 123 |
| GO:0007568 | aging | 2.03e-01 | 1.00e+00 | 2.145 | 1 | 2 | 123 |
| GO:0007050 | cell cycle arrest | 2.08e-01 | 1.00e+00 | 2.110 | 1 | 2 | 126 |
| GO:0007596 | blood coagulation | 2.09e-01 | 1.00e+00 | 1.230 | 2 | 5 | 464 |
| GO:0006511 | ubiquitin-dependent protein catabolic process | 2.09e-01 | 1.00e+00 | 2.099 | 1 | 3 | 127 |
| GO:0008201 | heparin binding | 2.09e-01 | 1.00e+00 | 2.099 | 1 | 1 | 127 |
| GO:0046983 | protein dimerization activity | 2.15e-01 | 1.00e+00 | 2.054 | 1 | 3 | 131 |
| GO:0006413 | translational initiation | 2.15e-01 | 1.00e+00 | 2.054 | 1 | 27 | 131 |
| GO:0042802 | identical protein binding | 2.28e-01 | 1.00e+00 | 1.148 | 2 | 4 | 491 |
| GO:0007507 | heart development | 2.29e-01 | 1.00e+00 | 1.948 | 1 | 1 | 141 |
| GO:0003735 | structural constituent of ribosome | 2.29e-01 | 1.00e+00 | 1.948 | 1 | 24 | 141 |
| GO:0005911 | cell-cell junction | 2.31e-01 | 1.00e+00 | 1.938 | 1 | 2 | 142 |
| GO:0008286 | insulin receptor signaling pathway | 2.34e-01 | 1.00e+00 | 1.918 | 1 | 4 | 144 |
| GO:0010628 | positive regulation of gene expression | 2.41e-01 | 1.00e+00 | 1.868 | 1 | 4 | 149 |
| GO:0008017 | microtubule binding | 2.42e-01 | 1.00e+00 | 1.859 | 1 | 2 | 150 |
| GO:0001666 | response to hypoxia | 2.42e-01 | 1.00e+00 | 1.859 | 1 | 1 | 150 |
| GO:0051260 | protein homooligomerization | 2.42e-01 | 1.00e+00 | 1.859 | 1 | 2 | 150 |
| GO:0000082 | G1/S transition of mitotic cell cycle | 2.42e-01 | 1.00e+00 | 1.859 | 1 | 11 | 150 |
| GO:0030246 | carbohydrate binding | 2.44e-01 | 1.00e+00 | 1.849 | 1 | 1 | 151 |
| GO:0042981 | regulation of apoptotic process | 2.44e-01 | 1.00e+00 | 1.849 | 1 | 7 | 151 |
| GO:0001077 | RNA polymerase II core promoter proximal region sequence-specific DNA binding transcription factor activity involved in positive regulation of transcription | 2.44e-01 | 1.00e+00 | 1.849 | 1 | 2 | 151 |
| GO:0043005 | neuron projection | 2.57e-01 | 1.00e+00 | 1.757 | 1 | 6 | 161 |
| GO:0016032 | viral process | 2.61e-01 | 1.00e+00 | 1.011 | 2 | 37 | 540 |
| GO:0000398 | mRNA splicing, via spliceosome | 2.63e-01 | 1.00e+00 | 1.721 | 1 | 2 | 165 |
| GO:0007601 | visual perception | 2.67e-01 | 1.00e+00 | 1.695 | 1 | 1 | 168 |
| GO:0016607 | nuclear speck | 2.77e-01 | 1.00e+00 | 1.636 | 1 | 2 | 175 |
| GO:0031965 | nuclear membrane | 2.78e-01 | 1.00e+00 | 1.628 | 1 | 2 | 176 |
| GO:0007049 | cell cycle | 2.79e-01 | 1.00e+00 | 1.620 | 1 | 4 | 177 |
| GO:0043687 | post-translational protein modification | 2.85e-01 | 1.00e+00 | 1.588 | 1 | 1 | 181 |
| GO:0015629 | actin cytoskeleton | 2.87e-01 | 1.00e+00 | 1.572 | 1 | 3 | 183 |
| GO:0005509 | calcium ion binding | 2.95e-01 | 1.00e+00 | 0.885 | 2 | 4 | 589 |
| GO:0007173 | epidermal growth factor receptor signaling pathway | 2.98e-01 | 1.00e+00 | 1.510 | 1 | 5 | 191 |
| GO:0045087 | innate immune response | 3.14e-01 | 1.00e+00 | 0.821 | 2 | 11 | 616 |
| GO:0042803 | protein homodimerization activity | 3.14e-01 | 1.00e+00 | 0.818 | 2 | 4 | 617 |
| GO:0030168 | platelet activation | 3.16e-01 | 1.00e+00 | 1.408 | 1 | 4 | 205 |
| GO:0001701 | in utero embryonic development | 3.22e-01 | 1.00e+00 | 1.373 | 1 | 2 | 210 |
| GO:0006351 | transcription, DNA-templated | 3.32e-01 | 1.00e+00 | 0.457 | 4 | 17 | 1585 |
| GO:0003713 | transcription coactivator activity | 3.58e-01 | 1.00e+00 | 1.187 | 1 | 6 | 239 |
| GO:0007399 | nervous system development | 3.65e-01 | 1.00e+00 | 1.151 | 1 | 1 | 245 |
| GO:0005730 | nucleolus | 3.75e-01 | 1.00e+00 | 0.370 | 4 | 36 | 1684 |
| GO:0005634 | nucleus | 3.91e-01 | 1.00e+00 | 0.172 | 10 | 66 | 4828 |
| GO:0043065 | positive regulation of apoptotic process | 3.99e-01 | 1.00e+00 | 0.990 | 1 | 6 | 274 |
| GO:0003779 | actin binding | 4.00e-01 | 1.00e+00 | 0.984 | 1 | 3 | 275 |
| GO:0007283 | spermatogenesis | 4.01e-01 | 1.00e+00 | 0.979 | 1 | 2 | 276 |
| GO:0030198 | extracellular matrix organization | 4.22e-01 | 1.00e+00 | 0.883 | 1 | 1 | 295 |
| GO:0030154 | cell differentiation | 4.53e-01 | 1.00e+00 | 0.743 | 1 | 3 | 325 |
| GO:0003677 | DNA binding | 4.57e-01 | 1.00e+00 | 0.273 | 3 | 14 | 1351 |
| GO:0007268 | synaptic transmission | 4.81e-01 | 1.00e+00 | 0.624 | 1 | 1 | 353 |
| GO:0009986 | cell surface | 5.45e-01 | 1.00e+00 | 0.366 | 1 | 1 | 422 |
| GO:0045892 | negative regulation of transcription, DNA-templated | 5.46e-01 | 1.00e+00 | 0.360 | 1 | 2 | 424 |
| GO:0006366 | transcription from RNA polymerase II promoter | 5.47e-01 | 1.00e+00 | 0.356 | 1 | 3 | 425 |
| GO:0005615 | extracellular space | 5.62e-01 | 1.00e+00 | 0.107 | 2 | 3 | 1010 |
| GO:0006468 | protein phosphorylation | 5.82e-01 | 1.00e+00 | 0.220 | 1 | 6 | 467 |
| GO:0055114 | oxidation-reduction process | 5.93e-01 | 1.00e+00 | 0.178 | 1 | 2 | 481 |
| GO:0045893 | positive regulation of transcription, DNA-templated | 5.97e-01 | 1.00e+00 | 0.160 | 1 | 8 | 487 |
| GO:0005654 | nucleoplasm | 6.07e-01 | 1.00e+00 | -0.009 | 2 | 26 | 1095 |
| GO:0006355 | regulation of transcription, DNA-templated | 6.12e-01 | 1.00e+00 | -0.021 | 2 | 10 | 1104 |
| GO:0000122 | negative regulation of transcription from RNA polymerase II promoter | 6.68e-01 | 1.00e+00 | -0.115 | 1 | 9 | 589 |
| GO:0005789 | endoplasmic reticulum membrane | 6.97e-01 | 1.00e+00 | -0.225 | 1 | 3 | 636 |
| GO:0010467 | gene expression | 7.15e-01 | 1.00e+00 | -0.298 | 1 | 36 | 669 |
| GO:0007165 | signal transduction | 8.35e-01 | 1.00e+00 | -0.804 | 1 | 7 | 950 |
| GO:0005887 | integral component of plasma membrane | 8.38e-01 | 1.00e+00 | -0.821 | 1 | 2 | 961 |
| GO:0008270 | zinc ion binding | 8.69e-01 | 1.00e+00 | -0.972 | 1 | 7 | 1067 |
| GO:0005524 | ATP binding | 9.26e-01 | 1.00e+00 | -1.315 | 1 | 19 | 1354 |
| GO:0046872 | metal ion binding | 9.41e-01 | 1.00e+00 | -1.429 | 1 | 14 | 1465 |
| GO:0016021 | integral component of membrane | 9.93e-01 | 1.00e+00 | -2.190 | 1 | 4 | 2483 |