meta-reg-snw-10097
Interacting subnets
| Subnetwork | Dataset | Score | p-value 1 | p-value 2 | p-value 3 | Size | Highlighted genes |
|---|---|---|---|---|---|---|---|
| int-snw-51741 | tai-screen-luciferase | 6.174 | 2.67e-126 | 1.32e-06 | 4.54e-03 | 24 | 19 |
| reg-snw-10097 | tai-screen-luciferase | 4.851 | 3.99e-26 | 2.69e-04 | 4.60e-04 | 6 | 3 |
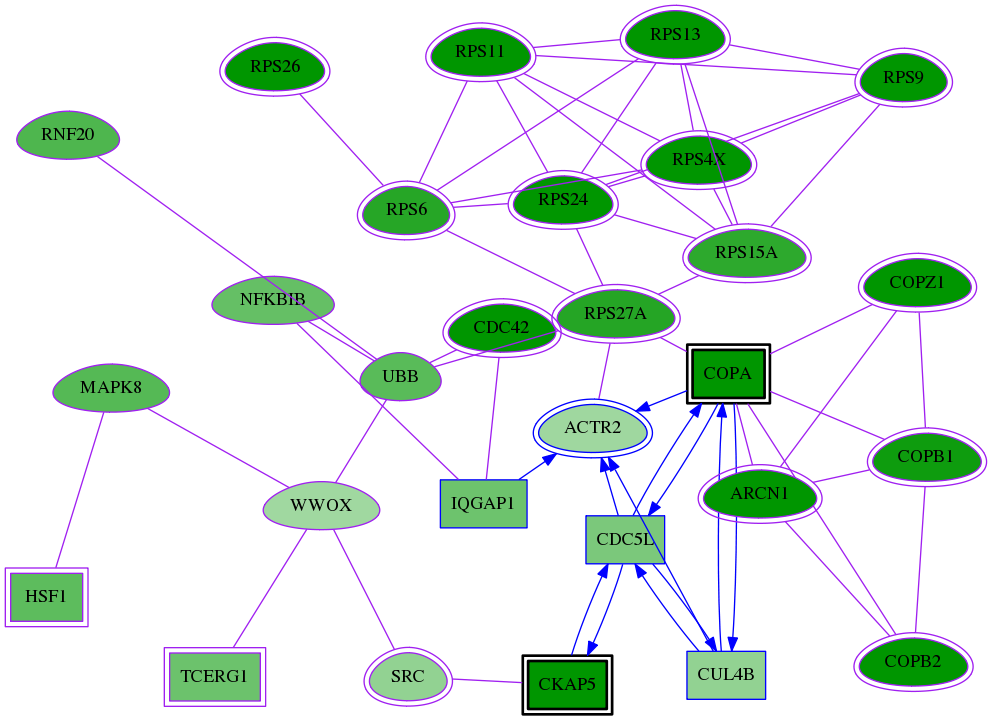
Genes (28)
| Gene Symbol | Entrez Gene ID | Frequency | tai-screen-luciferase gene score | Best subnetwork score | Degree | List-Gonzales GI | Tai-Hits |
|---|---|---|---|---|---|---|---|
| MAPK8 | 5599 | 28 | -4.403 | 6.468 | 153 | - | - |
| COPA | 1314 | 48 | -9.395 | 5.672 | 340 | Yes | Yes |
| RNF20 | 56254 | 28 | -4.564 | 6.174 | 18 | - | - |
| IQGAP1 | 8826 | 2 | -3.777 | 4.851 | 10 | - | - |
| CUL4B | 8450 | 21 | -2.809 | 5.269 | 142 | - | - |
| HSF1 | 3297 | 46 | -4.179 | 5.027 | 209 | - | Yes |
| RPS24 | 6229 | 46 | -7.034 | 8.389 | 217 | Yes | - |
| TCERG1 | 10915 | 28 | -3.808 | 6.174 | 58 | Yes | - |
| SRC | 6714 | 28 | -2.806 | 6.174 | 419 | Yes | - |
| RPS9 | 6203 | 45 | -7.127 | 7.555 | 140 | Yes | - |
| COPB2 | 9276 | 48 | -13.168 | 9.063 | 41 | Yes | Yes |
| CKAP5 | 9793 | 46 | -7.214 | 5.672 | 130 | Yes | Yes |
| RPS4X | 6191 | 44 | -6.747 | 7.555 | 263 | Yes | - |
| COPZ1 | 22818 | 48 | -8.301 | 9.063 | 13 | Yes | Yes |
| RPS15A | 6210 | 36 | -5.413 | 7.555 | 177 | Yes | - |
| CDC42 | 998 | 44 | -6.960 | 4.707 | 276 | Yes | Yes |
| WWOX | 51741 | 28 | -2.448 | 6.174 | 38 | - | - |
| RPS27A | 6233 | 45 | -5.631 | 8.389 | 344 | Yes | - |
| RPS11 | 6205 | 44 | -6.588 | 7.555 | 175 | Yes | - |
| RPS6 | 6194 | 44 | -5.603 | 8.046 | 217 | Yes | - |
| ACTR2 | 10097 | 2 | -2.490 | 4.851 | 65 | Yes | - |
| NFKBIB | 4793 | 28 | -3.978 | 6.174 | 78 | - | - |
| UBB | 7314 | 30 | -4.289 | 6.428 | 147 | - | - |
| RPS26 | 6231 | 43 | -7.478 | 8.046 | 60 | Yes | - |
| ARCN1 | 372 | 48 | -8.232 | 9.063 | 118 | Yes | Yes |
| COPB1 | 1315 | 39 | -6.221 | 9.063 | 118 | Yes | Yes |
| RPS13 | 6207 | 43 | -6.589 | 7.555 | 174 | Yes | - |
| CDC5L | 988 | 34 | -3.419 | 5.672 | 155 | - | - |
Interactions (55)
| Gene Symbol 1 | Entrez Gene ID 1 | Gene Symbol 2 | Entrez Gene ID 2 | Type | Direction | Origin databases / Sources |
|---|---|---|---|---|---|---|
| CDC42 | 998 | RPS27A | 6233 | pp | -- | int.I2D: SOURAV_MAPK_HIGH |
| ARCN1 | 372 | COPZ1 | 22818 | pp | -- | int.I2D: BioGrid_Yeast, IntAct_Yeast, MINT_Yeast, Tarassov_PCA, YeastMedium, INTEROLOG |
| RPS11 | 6205 | RPS15A | 6210 | pp | -- | int.I2D: BioGrid_Yeast |
| TCERG1 | 10915 | WWOX | 51741 | pp | -- | int.I2D: BioGrid, MINT, Pawson1; int.Mint: MI:0915(physical association) |
| RPS6 | 6194 | RPS13 | 6207 | pp | -- | int.I2D: BioGrid_Yeast |
| CDC5L | 988 | CUL4B | 8450 | pd | <> | reg.ITFP.txt: no annot |
| RPS11 | 6205 | RPS13 | 6207 | pp | -- | int.I2D: BioGrid_Yeast |
| SRC | 6714 | CKAP5 | 9793 | pp | -- | int.Intact: MI:0915(physical association); int.I2D: IntAct |
| RPS6 | 6194 | RPS26 | 6231 | pp | -- | int.I2D: YeastLow |
| CDC42 | 998 | UBB | 7314 | pp | -- | int.I2D: SOURAV_MAPK_HIGH |
| NFKBIB | 4793 | UBB | 7314 | pp | -- | int.Intact: MI:0915(physical association); int.I2D: IntAct |
| RPS9 | 6203 | RPS13 | 6207 | pp | -- | int.I2D: BioGrid_Yeast |
| RPS4X | 6191 | RPS11 | 6205 | pp | -- | int.I2D: BioGrid_Yeast, INTEROLOG, YeastMedium |
| RPS9 | 6203 | RPS24 | 6229 | pp | -- | int.I2D: BioGrid_Yeast |
| RPS27A | 6233 | ACTR2 | 10097 | pp | -- | int.I2D: YeastLow |
| RPS9 | 6203 | RPS15A | 6210 | pp | -- | int.I2D: BioGrid_Yeast |
| RPS6 | 6194 | RPS24 | 6229 | pp | -- | int.I2D: BioGrid_Yeast |
| RPS4X | 6191 | RPS9 | 6203 | pp | -- | int.I2D: BioGrid_Yeast |
| UBB | 7314 | WWOX | 51741 | pp | -- | int.I2D: BCI, HPRD; int.HPRD: in vivo |
| UBB | 7314 | RNF20 | 56254 | pp | -- | int.Intact: MI:0220(ubiquitination reaction); int.I2D: IntAct |
| RPS11 | 6205 | RPS24 | 6229 | pp | -- | int.I2D: BioGrid_Yeast |
| COPA | 1314 | ACTR2 | 10097 | pd | > | reg.ITFP.txt: no annot |
| NFKBIB | 4793 | IQGAP1 | 8826 | pp | -- | int.Intact: MI:0915(physical association); int.I2D: IntAct |
| SRC | 6714 | WWOX | 51741 | pp | -- | int.I2D: HPRD; int.HPRD: in vitro, in vivo |
| RPS27A | 6233 | UBB | 7314 | pp | -- | int.I2D: BIND |
| IQGAP1 | 8826 | ACTR2 | 10097 | pd | > | reg.ITFP.txt: no annot |
| RPS4X | 6191 | RPS24 | 6229 | pp | -- | int.I2D: BioGrid_Yeast |
| CUL4B | 8450 | ACTR2 | 10097 | pd | > | reg.ITFP.txt: no annot |
| RPS4X | 6191 | RPS15A | 6210 | pp | -- | int.I2D: BioGrid_Yeast, YeastMedium, INTEROLOG |
| RPS6 | 6194 | RPS11 | 6205 | pp | -- | int.I2D: BioGrid_Yeast |
| RPS13 | 6207 | RPS24 | 6229 | pp | -- | int.I2D: BioGrid_Yeast |
| RPS6 | 6194 | RPS27A | 6233 | pp | -- | int.I2D: YeastLow |
| COPA | 1314 | COPB1 | 1315 | pp | -- | int.I2D: BioGrid, BioGrid_Yeast, BIND_Yeast, IntAct_Yeast, MINT_Yeast, Krogan_Core, YeastHigh |
| CDC5L | 988 | CKAP5 | 9793 | pd | <> | reg.ITFP.txt: no annot |
| HSF1 | 3297 | MAPK8 | 5599 | pp | -- | int.I2D: HPRD, INNATEDB; int.HPRD: in vitro, in vivo |
| COPA | 1314 | COPZ1 | 22818 | pp | -- | int.I2D: BioGrid_Yeast, IntAct_Yeast, Krogan_Core, MINT_Yeast, YeastMedium |
| COPA | 1314 | COPB2 | 9276 | pp | -- | int.I2D: BioGrid, HPRD, BCI; int.HPRD: in vivo |
| COPB1 | 1315 | COPZ1 | 22818 | pp | -- | int.I2D: BioGrid_Yeast, IntAct_Yeast, Krogan_Core, MINT_Yeast, Tarassov_PCA, YeastMedium; int.DIP: MI:0915(physical association) |
| RPS4X | 6191 | RPS13 | 6207 | pp | -- | int.I2D: BioGrid_Yeast |
| RPS15A | 6210 | RPS24 | 6229 | pp | -- | int.I2D: BioGrid_Yeast |
| RPS13 | 6207 | RPS15A | 6210 | pp | -- | int.I2D: BioGrid_Yeast |
| CDC5L | 988 | COPA | 1314 | pd | <> | reg.ITFP.txt: no annot |
| COPA | 1314 | CUL4B | 8450 | pd | <> | reg.ITFP.txt: no annot |
| COPB1 | 1315 | COPB2 | 9276 | pp | -- | int.I2D: BioGrid |
| ARCN1 | 372 | COPB1 | 1315 | pp | -- | int.I2D: BCI, BioGrid, BioGrid_Yeast, IntAct_Yeast, INTEROLOG, MINT_Yeast, YeastHigh, BIND, BIND_Yeast, HPRD, Krogan_Core, MIPS; int.HPRD: in vivo |
| RPS9 | 6203 | RPS11 | 6205 | pp | -- | int.I2D: BioGrid_Yeast |
| CDC42 | 998 | IQGAP1 | 8826 | pp | -- | int.Intact: MI:0915(physical association); int.I2D: BioGrid, HPRD, MINT, BCI, IntAct; int.Mint: MI:0915(physical association), MI:0407(direct interaction); int.HPRD: in vitro, in vivo |
| ARCN1 | 372 | COPA | 1314 | pp | -- | int.I2D: BCI, BioGrid_Yeast, IntAct_Yeast, MINT_Yeast, YeastHigh, HPRD, Krogan_Core; int.HPRD: in vivo |
| RPS4X | 6191 | RPS6 | 6194 | pp | -- | int.I2D: BioGrid_Yeast |
| COPA | 1314 | RPS27A | 6233 | pp | -- | int.I2D: YeastLow |
| MAPK8 | 5599 | WWOX | 51741 | pp | -- | int.I2D: MINT, HPRD; int.Mint: MI:0915(physical association); int.HPRD: in vivo, yeast 2-hybrid |
| RPS15A | 6210 | RPS27A | 6233 | pp | -- | int.I2D: YeastLow |
| ARCN1 | 372 | COPB2 | 9276 | pp | -- | int.I2D: HPRD, BCI; int.HPRD: in vivo |
| CDC5L | 988 | ACTR2 | 10097 | pd | > | reg.ITFP.txt: no annot |
| RPS24 | 6229 | RPS27A | 6233 | pp | -- | int.I2D: YeastMedium |
Related GO terms (477)
| Accession number | Name | Hypergeometric test | Corrected p-value | Enrichment ratio | Occurrence in subnetwork | Occurrences in all snw genes | Occurrences in all int/reg genes |
|---|---|---|---|---|---|---|---|
| GO:0022627 | cytosolic small ribosomal subunit | 6.28e-18 | 1.02e-13 | 7.072 | 9 | 21 | 39 |
| GO:0019058 | viral life cycle | 2.39e-15 | 3.90e-11 | 5.664 | 10 | 25 | 115 |
| GO:0019083 | viral transcription | 7.39e-15 | 1.21e-10 | 6.017 | 9 | 22 | 81 |
| GO:0006415 | translational termination | 1.44e-14 | 2.36e-10 | 5.914 | 9 | 22 | 87 |
| GO:0006414 | translational elongation | 2.69e-14 | 4.39e-10 | 5.818 | 9 | 22 | 93 |
| GO:0006614 | SRP-dependent cotranslational protein targeting to membrane | 7.60e-14 | 1.24e-09 | 5.657 | 9 | 22 | 104 |
| GO:0000184 | nuclear-transcribed mRNA catabolic process, nonsense-mediated decay | 1.77e-13 | 2.90e-09 | 5.524 | 9 | 22 | 114 |
| GO:0005829 | cytosol | 2.70e-13 | 4.40e-09 | 2.323 | 22 | 74 | 2562 |
| GO:0006413 | translational initiation | 6.36e-13 | 1.04e-08 | 5.324 | 9 | 27 | 131 |
| GO:0003735 | structural constituent of ribosome | 1.24e-12 | 2.03e-08 | 5.217 | 9 | 24 | 141 |
| GO:0016071 | mRNA metabolic process | 1.96e-12 | 3.20e-08 | 4.708 | 10 | 29 | 223 |
| GO:0016070 | RNA metabolic process | 5.43e-12 | 8.86e-08 | 4.561 | 10 | 29 | 247 |
| GO:0048205 | COPI coating of Golgi vesicle | 1.30e-11 | 2.12e-07 | 7.809 | 5 | 6 | 13 |
| GO:0030126 | COPI vesicle coat | 1.30e-11 | 2.12e-07 | 7.809 | 5 | 6 | 13 |
| GO:0016032 | viral process | 2.86e-11 | 4.67e-07 | 3.695 | 12 | 37 | 540 |
| GO:0015935 | small ribosomal subunit | 6.22e-11 | 1.01e-06 | 7.422 | 5 | 9 | 17 |
| GO:0061024 | membrane organization | 9.03e-11 | 1.47e-06 | 4.997 | 8 | 11 | 146 |
| GO:0006412 | translation | 1.24e-10 | 2.03e-06 | 4.480 | 9 | 29 | 235 |
| GO:0006890 | retrograde vesicle-mediated transport, Golgi to ER | 5.29e-10 | 8.63e-06 | 6.865 | 5 | 6 | 25 |
| GO:0016020 | membrane | 2.39e-09 | 3.90e-05 | 2.417 | 16 | 48 | 1746 |
| GO:0036464 | cytoplasmic ribonucleoprotein granule | 4.96e-08 | 8.10e-04 | 6.728 | 4 | 5 | 22 |
| GO:0044822 | poly(A) RNA binding | 7.34e-08 | 1.20e-03 | 2.698 | 12 | 42 | 1078 |
| GO:0044267 | cellular protein metabolic process | 8.34e-08 | 1.36e-03 | 3.406 | 9 | 29 | 495 |
| GO:0010467 | gene expression | 8.41e-08 | 1.37e-03 | 3.123 | 10 | 36 | 669 |
| GO:0005925 | focal adhesion | 2.28e-06 | 3.73e-02 | 3.463 | 7 | 23 | 370 |
| GO:0005840 | ribosome | 2.96e-06 | 4.82e-02 | 5.304 | 4 | 10 | 59 |
| GO:0006891 | intra-Golgi vesicle-mediated transport | 3.03e-06 | 4.94e-02 | 6.685 | 3 | 3 | 17 |
| GO:0034166 | toll-like receptor 10 signaling pathway | 4.37e-06 | 7.12e-02 | 5.165 | 4 | 5 | 65 |
| GO:0034146 | toll-like receptor 5 signaling pathway | 4.37e-06 | 7.12e-02 | 5.165 | 4 | 5 | 65 |
| GO:0038123 | toll-like receptor TLR1:TLR2 signaling pathway | 6.22e-06 | 1.02e-01 | 5.037 | 4 | 5 | 71 |
| GO:0038124 | toll-like receptor TLR6:TLR2 signaling pathway | 6.22e-06 | 1.02e-01 | 5.037 | 4 | 5 | 71 |
| GO:0034162 | toll-like receptor 9 signaling pathway | 6.58e-06 | 1.07e-01 | 5.017 | 4 | 5 | 72 |
| GO:0034134 | toll-like receptor 2 signaling pathway | 6.95e-06 | 1.13e-01 | 4.997 | 4 | 5 | 73 |
| GO:0035666 | TRIF-dependent toll-like receptor signaling pathway | 8.17e-06 | 1.33e-01 | 4.939 | 4 | 5 | 76 |
| GO:0002756 | MyD88-independent toll-like receptor signaling pathway | 9.06e-06 | 1.48e-01 | 4.902 | 4 | 5 | 78 |
| GO:0002755 | MyD88-dependent toll-like receptor signaling pathway | 1.00e-05 | 1.64e-01 | 4.865 | 4 | 5 | 80 |
| GO:0034138 | toll-like receptor 3 signaling pathway | 1.00e-05 | 1.64e-01 | 4.865 | 4 | 5 | 80 |
| GO:0006886 | intracellular protein transport | 1.02e-05 | 1.66e-01 | 4.074 | 5 | 6 | 173 |
| GO:0019843 | rRNA binding | 1.29e-05 | 2.10e-01 | 6.017 | 3 | 4 | 27 |
| GO:0007173 | epidermal growth factor receptor signaling pathway | 1.64e-05 | 2.68e-01 | 3.932 | 5 | 5 | 191 |
| GO:0070062 | extracellular vesicular exosome | 1.94e-05 | 3.16e-01 | 1.698 | 14 | 51 | 2516 |
| GO:0034142 | toll-like receptor 4 signaling pathway | 2.07e-05 | 3.37e-01 | 4.602 | 4 | 5 | 96 |
| GO:0033119 | negative regulation of RNA splicing | 2.83e-05 | 4.62e-01 | 7.865 | 2 | 2 | 5 |
| GO:0042059 | negative regulation of epidermal growth factor receptor signaling pathway | 3.11e-05 | 5.07e-01 | 5.602 | 3 | 3 | 36 |
| GO:0002224 | toll-like receptor signaling pathway | 3.41e-05 | 5.56e-01 | 4.419 | 4 | 5 | 109 |
| GO:0045087 | innate immune response | 6.25e-05 | 1.00e+00 | 2.728 | 7 | 11 | 616 |
| GO:0007254 | JNK cascade | 8.91e-05 | 1.00e+00 | 5.100 | 3 | 3 | 51 |
| GO:0043065 | positive regulation of apoptotic process | 9.21e-05 | 1.00e+00 | 3.411 | 5 | 6 | 274 |
| GO:0051403 | stress-activated MAPK cascade | 1.06e-04 | 1.00e+00 | 5.017 | 3 | 4 | 54 |
| GO:0032481 | positive regulation of type I interferon production | 1.52e-04 | 1.00e+00 | 4.841 | 3 | 3 | 61 |
| GO:0043005 | neuron projection | 1.55e-04 | 1.00e+00 | 3.856 | 4 | 6 | 161 |
| GO:0042274 | ribosomal small subunit biogenesis | 1.85e-04 | 1.00e+00 | 6.602 | 2 | 6 | 12 |
| GO:0019082 | viral protein processing | 1.85e-04 | 1.00e+00 | 6.602 | 2 | 2 | 12 |
| GO:0005737 | cytoplasm | 2.08e-04 | 1.00e+00 | 1.230 | 16 | 65 | 3976 |
| GO:0032479 | regulation of type I interferon production | 2.19e-04 | 1.00e+00 | 6.487 | 2 | 2 | 13 |
| GO:0038096 | Fc-gamma receptor signaling pathway involved in phagocytosis | 3.04e-04 | 1.00e+00 | 4.505 | 3 | 4 | 77 |
| GO:0005730 | nucleolus | 3.05e-04 | 1.00e+00 | 1.791 | 10 | 36 | 1684 |
| GO:0075733 | intracellular transport of virus | 3.80e-04 | 1.00e+00 | 6.100 | 2 | 3 | 17 |
| GO:0019068 | virion assembly | 3.80e-04 | 1.00e+00 | 6.100 | 2 | 2 | 17 |
| GO:0071456 | cellular response to hypoxia | 6.17e-04 | 1.00e+00 | 4.157 | 3 | 3 | 98 |
| GO:0007220 | Notch receptor processing | 6.42e-04 | 1.00e+00 | 5.728 | 2 | 2 | 22 |
| GO:0043066 | negative regulation of apoptotic process | 7.62e-04 | 1.00e+00 | 2.751 | 5 | 14 | 433 |
| GO:0007163 | establishment or maintenance of cell polarity | 7.65e-04 | 1.00e+00 | 5.602 | 2 | 2 | 24 |
| GO:0070423 | nucleotide-binding oligomerization domain containing signaling pathway | 8.31e-04 | 1.00e+00 | 5.543 | 2 | 2 | 25 |
| GO:0005844 | polysome | 8.31e-04 | 1.00e+00 | 5.543 | 2 | 2 | 25 |
| GO:0005978 | glycogen biosynthetic process | 8.99e-04 | 1.00e+00 | 5.487 | 2 | 2 | 26 |
| GO:0061418 | regulation of transcription from RNA polymerase II promoter in response to hypoxia | 8.99e-04 | 1.00e+00 | 5.487 | 2 | 2 | 26 |
| GO:0043552 | positive regulation of phosphatidylinositol 3-kinase activity | 8.99e-04 | 1.00e+00 | 5.487 | 2 | 2 | 26 |
| GO:0030529 | ribonucleoprotein complex | 9.57e-04 | 1.00e+00 | 3.939 | 3 | 8 | 114 |
| GO:0097190 | apoptotic signaling pathway | 1.01e-03 | 1.00e+00 | 3.914 | 3 | 3 | 116 |
| GO:0000209 | protein polyubiquitination | 1.01e-03 | 1.00e+00 | 3.914 | 3 | 7 | 116 |
| GO:0005515 | protein binding | 1.10e-03 | 1.00e+00 | 0.854 | 19 | 87 | 6127 |
| GO:0048011 | neurotrophin TRK receptor signaling pathway | 1.14e-03 | 1.00e+00 | 3.094 | 4 | 5 | 273 |
| GO:0051092 | positive regulation of NF-kappaB transcription factor activity | 1.25e-03 | 1.00e+00 | 3.806 | 3 | 3 | 125 |
| GO:0032480 | negative regulation of type I interferon production | 1.36e-03 | 1.00e+00 | 5.187 | 2 | 2 | 32 |
| GO:0007179 | transforming growth factor beta receptor signaling pathway | 1.40e-03 | 1.00e+00 | 3.750 | 3 | 4 | 130 |
| GO:0000086 | G2/M transition of mitotic cell cycle | 1.63e-03 | 1.00e+00 | 3.674 | 3 | 6 | 137 |
| GO:2001286 | regulation of caveolin-mediated endocytosis | 1.72e-03 | 1.00e+00 | 9.187 | 1 | 1 | 1 |
| GO:0034332 | adherens junction organization | 1.72e-03 | 1.00e+00 | 5.017 | 2 | 4 | 36 |
| GO:0022605 | oogenesis stage | 1.72e-03 | 1.00e+00 | 9.187 | 1 | 1 | 1 |
| GO:0071393 | cellular response to progesterone stimulus | 1.72e-03 | 1.00e+00 | 9.187 | 1 | 1 | 1 |
| GO:0021691 | cerebellar Purkinje cell layer maturation | 1.72e-03 | 1.00e+00 | 9.187 | 1 | 1 | 1 |
| GO:0000082 | G1/S transition of mitotic cell cycle | 2.11e-03 | 1.00e+00 | 3.543 | 3 | 11 | 150 |
| GO:0007249 | I-kappaB kinase/NF-kappaB signaling | 2.23e-03 | 1.00e+00 | 4.830 | 2 | 4 | 41 |
| GO:0035872 | nucleotide-binding domain, leucine rich repeat containing receptor signaling pathway | 2.45e-03 | 1.00e+00 | 4.761 | 2 | 2 | 43 |
| GO:0005198 | structural molecule activity | 2.49e-03 | 1.00e+00 | 3.459 | 3 | 5 | 159 |
| GO:0008543 | fibroblast growth factor receptor signaling pathway | 2.49e-03 | 1.00e+00 | 3.459 | 3 | 3 | 159 |
| GO:0038095 | Fc-epsilon receptor signaling pathway | 3.05e-03 | 1.00e+00 | 3.354 | 3 | 3 | 171 |
| GO:0071338 | positive regulation of hair follicle cell proliferation | 3.43e-03 | 1.00e+00 | 8.187 | 1 | 1 | 2 |
| GO:0033206 | meiotic cytokinesis | 3.43e-03 | 1.00e+00 | 8.187 | 1 | 1 | 2 |
| GO:0071987 | WD40-repeat domain binding | 3.43e-03 | 1.00e+00 | 8.187 | 1 | 1 | 2 |
| GO:0060661 | submandibular salivary gland formation | 3.43e-03 | 1.00e+00 | 8.187 | 1 | 1 | 2 |
| GO:0072422 | signal transduction involved in DNA damage checkpoint | 3.43e-03 | 1.00e+00 | 8.187 | 1 | 1 | 2 |
| GO:1902255 | positive regulation of intrinsic apoptotic signaling pathway by p53 class mediator | 3.43e-03 | 1.00e+00 | 8.187 | 1 | 1 | 2 |
| GO:0016344 | meiotic chromosome movement towards spindle pole | 3.43e-03 | 1.00e+00 | 8.187 | 1 | 1 | 2 |
| GO:0035305 | negative regulation of dephosphorylation | 3.43e-03 | 1.00e+00 | 8.187 | 1 | 1 | 2 |
| GO:0010632 | regulation of epithelial cell migration | 3.43e-03 | 1.00e+00 | 8.187 | 1 | 1 | 2 |
| GO:0090135 | actin filament branching | 3.43e-03 | 1.00e+00 | 8.187 | 1 | 1 | 2 |
| GO:0030666 | endocytic vesicle membrane | 3.84e-03 | 1.00e+00 | 4.432 | 2 | 2 | 54 |
| GO:0016197 | endosomal transport | 4.42e-03 | 1.00e+00 | 4.329 | 2 | 3 | 58 |
| GO:0051683 | establishment of Golgi localization | 5.14e-03 | 1.00e+00 | 7.602 | 1 | 2 | 3 |
| GO:0010641 | positive regulation of platelet-derived growth factor receptor signaling pathway | 5.14e-03 | 1.00e+00 | 7.602 | 1 | 1 | 3 |
| GO:0001162 | RNA polymerase II intronic transcription regulatory region sequence-specific DNA binding | 5.14e-03 | 1.00e+00 | 7.602 | 1 | 1 | 3 |
| GO:0008356 | asymmetric cell division | 5.14e-03 | 1.00e+00 | 7.602 | 1 | 1 | 3 |
| GO:0003161 | cardiac conduction system development | 5.14e-03 | 1.00e+00 | 7.602 | 1 | 1 | 3 |
| GO:0051653 | spindle localization | 5.14e-03 | 1.00e+00 | 7.602 | 1 | 1 | 3 |
| GO:0030951 | establishment or maintenance of microtubule cytoskeleton polarity | 5.14e-03 | 1.00e+00 | 7.602 | 1 | 1 | 3 |
| GO:0033146 | regulation of intracellular estrogen receptor signaling pathway | 5.14e-03 | 1.00e+00 | 7.602 | 1 | 1 | 3 |
| GO:0090045 | positive regulation of deacetylase activity | 5.14e-03 | 1.00e+00 | 7.602 | 1 | 1 | 3 |
| GO:0035033 | histone deacetylase regulator activity | 5.14e-03 | 1.00e+00 | 7.602 | 1 | 1 | 3 |
| GO:2000017 | positive regulation of determination of dorsal identity | 5.14e-03 | 1.00e+00 | 7.602 | 1 | 1 | 3 |
| GO:0004705 | JUN kinase activity | 5.14e-03 | 1.00e+00 | 7.602 | 1 | 1 | 3 |
| GO:0030478 | actin cap | 5.14e-03 | 1.00e+00 | 7.602 | 1 | 1 | 3 |
| GO:0030512 | negative regulation of transforming growth factor beta receptor signaling pathway | 5.36e-03 | 1.00e+00 | 4.187 | 2 | 2 | 64 |
| GO:0051436 | negative regulation of ubiquitin-protein ligase activity involved in mitotic cell cycle | 5.52e-03 | 1.00e+00 | 4.165 | 2 | 6 | 65 |
| GO:0006977 | DNA damage response, signal transduction by p53 class mediator resulting in cell cycle arrest | 5.52e-03 | 1.00e+00 | 4.165 | 2 | 6 | 65 |
| GO:0031295 | T cell costimulation | 5.86e-03 | 1.00e+00 | 4.121 | 2 | 3 | 67 |
| GO:0018105 | peptidyl-serine phosphorylation | 6.20e-03 | 1.00e+00 | 4.079 | 2 | 5 | 69 |
| GO:0051437 | positive regulation of ubiquitin-protein ligase activity involved in mitotic cell cycle | 6.38e-03 | 1.00e+00 | 4.058 | 2 | 8 | 70 |
| GO:0007258 | JUN phosphorylation | 6.85e-03 | 1.00e+00 | 7.187 | 1 | 1 | 4 |
| GO:0034191 | apolipoprotein A-I receptor binding | 6.85e-03 | 1.00e+00 | 7.187 | 1 | 1 | 4 |
| GO:0048664 | neuron fate determination | 6.85e-03 | 1.00e+00 | 7.187 | 1 | 1 | 4 |
| GO:0033503 | HULC complex | 6.85e-03 | 1.00e+00 | 7.187 | 1 | 1 | 4 |
| GO:0032463 | negative regulation of protein homooligomerization | 6.85e-03 | 1.00e+00 | 7.187 | 1 | 1 | 4 |
| GO:0051902 | negative regulation of mitochondrial depolarization | 6.85e-03 | 1.00e+00 | 7.187 | 1 | 1 | 4 |
| GO:0031062 | positive regulation of histone methylation | 6.85e-03 | 1.00e+00 | 7.187 | 1 | 1 | 4 |
| GO:0031063 | regulation of histone deacetylation | 6.85e-03 | 1.00e+00 | 7.187 | 1 | 1 | 4 |
| GO:0060684 | epithelial-mesenchymal cell signaling | 6.85e-03 | 1.00e+00 | 7.187 | 1 | 1 | 4 |
| GO:0070851 | growth factor receptor binding | 6.85e-03 | 1.00e+00 | 7.187 | 1 | 1 | 4 |
| GO:0051835 | positive regulation of synapse structural plasticity | 6.85e-03 | 1.00e+00 | 7.187 | 1 | 1 | 4 |
| GO:0033625 | positive regulation of integrin activation | 6.85e-03 | 1.00e+00 | 7.187 | 1 | 1 | 4 |
| GO:0090231 | regulation of spindle checkpoint | 6.85e-03 | 1.00e+00 | 7.187 | 1 | 1 | 4 |
| GO:0072384 | organelle transport along microtubule | 6.85e-03 | 1.00e+00 | 7.187 | 1 | 2 | 4 |
| GO:0002479 | antigen processing and presentation of exogenous peptide antigen via MHC class I, TAP-dependent | 6.92e-03 | 1.00e+00 | 3.997 | 2 | 6 | 73 |
| GO:0003729 | mRNA binding | 6.92e-03 | 1.00e+00 | 3.997 | 2 | 3 | 73 |
| GO:0051439 | regulation of ubiquitin-protein ligase activity involved in mitotic cell cycle | 7.10e-03 | 1.00e+00 | 3.978 | 2 | 8 | 74 |
| GO:0042590 | antigen processing and presentation of exogenous peptide antigen via MHC class I | 7.67e-03 | 1.00e+00 | 3.920 | 2 | 6 | 77 |
| GO:0003713 | transcription coactivator activity | 7.75e-03 | 1.00e+00 | 2.871 | 3 | 6 | 239 |
| GO:0031145 | anaphase-promoting complex-dependent proteasomal ubiquitin-dependent protein catabolic process | 8.06e-03 | 1.00e+00 | 3.883 | 2 | 8 | 79 |
| GO:0071222 | cellular response to lipopolysaccharide | 8.46e-03 | 1.00e+00 | 3.847 | 2 | 4 | 81 |
| GO:0031465 | Cul4B-RING E3 ubiquitin ligase complex | 8.55e-03 | 1.00e+00 | 6.865 | 1 | 1 | 5 |
| GO:1990138 | neuron projection extension | 8.55e-03 | 1.00e+00 | 6.865 | 1 | 1 | 5 |
| GO:2000641 | regulation of early endosome to late endosome transport | 8.55e-03 | 1.00e+00 | 6.865 | 1 | 1 | 5 |
| GO:0036336 | dendritic cell migration | 8.55e-03 | 1.00e+00 | 6.865 | 1 | 1 | 5 |
| GO:0035088 | establishment or maintenance of apical/basal cell polarity | 8.55e-03 | 1.00e+00 | 6.865 | 1 | 1 | 5 |
| GO:0051385 | response to mineralocorticoid | 8.55e-03 | 1.00e+00 | 6.865 | 1 | 1 | 5 |
| GO:0000462 | maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) | 8.55e-03 | 1.00e+00 | 6.865 | 1 | 3 | 5 |
| GO:0097300 | programmed necrotic cell death | 8.55e-03 | 1.00e+00 | 6.865 | 1 | 1 | 5 |
| GO:0031256 | leading edge membrane | 8.55e-03 | 1.00e+00 | 6.865 | 1 | 1 | 5 |
| GO:0071803 | positive regulation of podosome assembly | 8.55e-03 | 1.00e+00 | 6.865 | 1 | 1 | 5 |
| GO:0016482 | cytoplasmic transport | 8.55e-03 | 1.00e+00 | 6.865 | 1 | 1 | 5 |
| GO:0050852 | T cell receptor signaling pathway | 9.92e-03 | 1.00e+00 | 3.728 | 2 | 2 | 88 |
| GO:0050847 | progesterone receptor signaling pathway | 1.03e-02 | 1.00e+00 | 6.602 | 1 | 1 | 6 |
| GO:0007143 | female meiotic division | 1.03e-02 | 1.00e+00 | 6.602 | 1 | 1 | 6 |
| GO:0002309 | T cell proliferation involved in immune response | 1.03e-02 | 1.00e+00 | 6.602 | 1 | 1 | 6 |
| GO:0045182 | translation regulator activity | 1.03e-02 | 1.00e+00 | 6.602 | 1 | 2 | 6 |
| GO:0045056 | transcytosis | 1.03e-02 | 1.00e+00 | 6.602 | 1 | 1 | 6 |
| GO:0000974 | Prp19 complex | 1.03e-02 | 1.00e+00 | 6.602 | 1 | 1 | 6 |
| GO:0060789 | hair follicle placode formation | 1.03e-02 | 1.00e+00 | 6.602 | 1 | 1 | 6 |
| GO:0072015 | glomerular visceral epithelial cell development | 1.03e-02 | 1.00e+00 | 6.602 | 1 | 1 | 6 |
| GO:0048554 | positive regulation of metalloenzyme activity | 1.03e-02 | 1.00e+00 | 6.602 | 1 | 1 | 6 |
| GO:0006924 | activation-induced cell death of T cells | 1.03e-02 | 1.00e+00 | 6.602 | 1 | 2 | 6 |
| GO:0000187 | activation of MAPK activity | 1.04e-02 | 1.00e+00 | 3.695 | 2 | 3 | 90 |
| GO:0002474 | antigen processing and presentation of peptide antigen via MHC class I | 1.13e-02 | 1.00e+00 | 3.632 | 2 | 7 | 94 |
| GO:0001649 | osteoblast differentiation | 1.15e-02 | 1.00e+00 | 3.617 | 2 | 3 | 95 |
| GO:0006364 | rRNA processing | 1.17e-02 | 1.00e+00 | 3.602 | 2 | 8 | 96 |
| GO:0050658 | RNA transport | 1.19e-02 | 1.00e+00 | 6.380 | 1 | 1 | 7 |
| GO:0007097 | nuclear migration | 1.19e-02 | 1.00e+00 | 6.380 | 1 | 2 | 7 |
| GO:0003334 | keratinocyte development | 1.19e-02 | 1.00e+00 | 6.380 | 1 | 1 | 7 |
| GO:0051988 | regulation of attachment of spindle microtubules to kinetochore | 1.19e-02 | 1.00e+00 | 6.380 | 1 | 1 | 7 |
| GO:0070914 | UV-damage excision repair | 1.19e-02 | 1.00e+00 | 6.380 | 1 | 1 | 7 |
| GO:0000028 | ribosomal small subunit assembly | 1.19e-02 | 1.00e+00 | 6.380 | 1 | 3 | 7 |
| GO:0010447 | response to acidic pH | 1.19e-02 | 1.00e+00 | 6.380 | 1 | 1 | 7 |
| GO:0034101 | erythrocyte homeostasis | 1.19e-02 | 1.00e+00 | 6.380 | 1 | 1 | 7 |
| GO:0005885 | Arp2/3 protein complex | 1.19e-02 | 1.00e+00 | 6.380 | 1 | 1 | 7 |
| GO:0010907 | positive regulation of glucose metabolic process | 1.19e-02 | 1.00e+00 | 6.380 | 1 | 1 | 7 |
| GO:0030157 | pancreatic juice secretion | 1.19e-02 | 1.00e+00 | 6.380 | 1 | 1 | 7 |
| GO:0060136 | embryonic process involved in female pregnancy | 1.19e-02 | 1.00e+00 | 6.380 | 1 | 1 | 7 |
| GO:0000930 | gamma-tubulin complex | 1.19e-02 | 1.00e+00 | 6.380 | 1 | 1 | 7 |
| GO:0043497 | regulation of protein heterodimerization activity | 1.19e-02 | 1.00e+00 | 6.380 | 1 | 1 | 7 |
| GO:0043114 | regulation of vascular permeability | 1.36e-02 | 1.00e+00 | 6.187 | 1 | 1 | 8 |
| GO:0045124 | regulation of bone resorption | 1.36e-02 | 1.00e+00 | 6.187 | 1 | 1 | 8 |
| GO:0051489 | regulation of filopodium assembly | 1.36e-02 | 1.00e+00 | 6.187 | 1 | 1 | 8 |
| GO:0033523 | histone H2B ubiquitination | 1.36e-02 | 1.00e+00 | 6.187 | 1 | 1 | 8 |
| GO:0036057 | slit diaphragm | 1.36e-02 | 1.00e+00 | 6.187 | 1 | 1 | 8 |
| GO:0007172 | signal complex assembly | 1.36e-02 | 1.00e+00 | 6.187 | 1 | 1 | 8 |
| GO:0030496 | midbody | 1.49e-02 | 1.00e+00 | 3.419 | 2 | 4 | 109 |
| GO:0090136 | epithelial cell-cell adhesion | 1.53e-02 | 1.00e+00 | 6.017 | 1 | 2 | 9 |
| GO:0047497 | mitochondrion transport along microtubule | 1.53e-02 | 1.00e+00 | 6.017 | 1 | 1 | 9 |
| GO:0071732 | cellular response to nitric oxide | 1.53e-02 | 1.00e+00 | 6.017 | 1 | 1 | 9 |
| GO:0046628 | positive regulation of insulin receptor signaling pathway | 1.53e-02 | 1.00e+00 | 6.017 | 1 | 1 | 9 |
| GO:0015630 | microtubule cytoskeleton | 1.57e-02 | 1.00e+00 | 3.380 | 2 | 4 | 112 |
| GO:0031274 | positive regulation of pseudopodium assembly | 1.70e-02 | 1.00e+00 | 5.865 | 1 | 2 | 10 |
| GO:0019901 | protein kinase binding | 1.70e-02 | 1.00e+00 | 2.450 | 3 | 9 | 320 |
| GO:0022407 | regulation of cell-cell adhesion | 1.70e-02 | 1.00e+00 | 5.865 | 1 | 1 | 10 |
| GO:0001817 | regulation of cytokine production | 1.70e-02 | 1.00e+00 | 5.865 | 1 | 1 | 10 |
| GO:0005798 | Golgi-associated vesicle | 1.70e-02 | 1.00e+00 | 5.865 | 1 | 1 | 10 |
| GO:0060047 | heart contraction | 1.70e-02 | 1.00e+00 | 5.865 | 1 | 1 | 10 |
| GO:0000122 | negative regulation of transcription from RNA polymerase II promoter | 1.73e-02 | 1.00e+00 | 1.985 | 4 | 9 | 589 |
| GO:0006006 | glucose metabolic process | 1.76e-02 | 1.00e+00 | 3.292 | 2 | 4 | 119 |
| GO:0005634 | nucleus | 1.83e-02 | 1.00e+00 | 0.757 | 14 | 66 | 4828 |
| GO:0045737 | positive regulation of cyclin-dependent protein serine/threonine kinase activity | 1.87e-02 | 1.00e+00 | 5.728 | 1 | 1 | 11 |
| GO:0034314 | Arp2/3 complex-mediated actin nucleation | 1.87e-02 | 1.00e+00 | 5.728 | 1 | 1 | 11 |
| GO:0060065 | uterus development | 1.87e-02 | 1.00e+00 | 5.728 | 1 | 1 | 11 |
| GO:0051895 | negative regulation of focal adhesion assembly | 1.87e-02 | 1.00e+00 | 5.728 | 1 | 1 | 11 |
| GO:0045120 | pronucleus | 1.87e-02 | 1.00e+00 | 5.728 | 1 | 1 | 11 |
| GO:0010390 | histone monoubiquitination | 1.87e-02 | 1.00e+00 | 5.728 | 1 | 1 | 11 |
| GO:0035518 | histone H2A monoubiquitination | 1.87e-02 | 1.00e+00 | 5.728 | 1 | 2 | 11 |
| GO:2000573 | positive regulation of DNA biosynthetic process | 1.87e-02 | 1.00e+00 | 5.728 | 1 | 2 | 11 |
| GO:0003682 | chromatin binding | 1.91e-02 | 1.00e+00 | 2.388 | 3 | 4 | 334 |
| GO:0007219 | Notch signaling pathway | 1.93e-02 | 1.00e+00 | 3.221 | 2 | 4 | 125 |
| GO:0006511 | ubiquitin-dependent protein catabolic process | 1.99e-02 | 1.00e+00 | 3.198 | 2 | 3 | 127 |
| GO:0061136 | regulation of proteasomal protein catabolic process | 2.04e-02 | 1.00e+00 | 5.602 | 1 | 1 | 12 |
| GO:0007051 | spindle organization | 2.04e-02 | 1.00e+00 | 5.602 | 1 | 1 | 12 |
| GO:0043149 | stress fiber assembly | 2.04e-02 | 1.00e+00 | 5.602 | 1 | 2 | 12 |
| GO:1901214 | regulation of neuron death | 2.04e-02 | 1.00e+00 | 5.602 | 1 | 1 | 12 |
| GO:0009615 | response to virus | 2.14e-02 | 1.00e+00 | 3.143 | 2 | 4 | 132 |
| GO:0031929 | TOR signaling | 2.21e-02 | 1.00e+00 | 5.487 | 1 | 1 | 13 |
| GO:0005662 | DNA replication factor A complex | 2.21e-02 | 1.00e+00 | 5.487 | 1 | 1 | 13 |
| GO:0071398 | cellular response to fatty acid | 2.21e-02 | 1.00e+00 | 5.487 | 1 | 2 | 13 |
| GO:1900087 | positive regulation of G1/S transition of mitotic cell cycle | 2.21e-02 | 1.00e+00 | 5.487 | 1 | 1 | 13 |
| GO:0060444 | branching involved in mammary gland duct morphogenesis | 2.21e-02 | 1.00e+00 | 5.487 | 1 | 1 | 13 |
| GO:0035371 | microtubule plus-end | 2.38e-02 | 1.00e+00 | 5.380 | 1 | 1 | 14 |
| GO:0031333 | negative regulation of protein complex assembly | 2.38e-02 | 1.00e+00 | 5.380 | 1 | 1 | 14 |
| GO:0031996 | thioesterase binding | 2.38e-02 | 1.00e+00 | 5.380 | 1 | 1 | 14 |
| GO:0050662 | coenzyme binding | 2.38e-02 | 1.00e+00 | 5.380 | 1 | 1 | 14 |
| GO:0005095 | GTPase inhibitor activity | 2.38e-02 | 1.00e+00 | 5.380 | 1 | 1 | 14 |
| GO:0043539 | protein serine/threonine kinase activator activity | 2.38e-02 | 1.00e+00 | 5.380 | 1 | 1 | 14 |
| GO:0048821 | erythrocyte development | 2.54e-02 | 1.00e+00 | 5.280 | 1 | 2 | 15 |
| GO:2001241 | positive regulation of extrinsic apoptotic signaling pathway in absence of ligand | 2.54e-02 | 1.00e+00 | 5.280 | 1 | 1 | 15 |
| GO:0048477 | oogenesis | 2.54e-02 | 1.00e+00 | 5.280 | 1 | 1 | 15 |
| GO:0030131 | clathrin adaptor complex | 2.54e-02 | 1.00e+00 | 5.280 | 1 | 1 | 15 |
| GO:0030225 | macrophage differentiation | 2.54e-02 | 1.00e+00 | 5.280 | 1 | 1 | 15 |
| GO:0031369 | translation initiation factor binding | 2.54e-02 | 1.00e+00 | 5.280 | 1 | 2 | 15 |
| GO:0051233 | spindle midzone | 2.54e-02 | 1.00e+00 | 5.280 | 1 | 2 | 15 |
| GO:0043123 | positive regulation of I-kappaB kinase/NF-kappaB signaling | 2.65e-02 | 1.00e+00 | 2.978 | 2 | 3 | 148 |
| GO:0010628 | positive regulation of gene expression | 2.68e-02 | 1.00e+00 | 2.968 | 2 | 4 | 149 |
| GO:0042176 | regulation of protein catabolic process | 2.71e-02 | 1.00e+00 | 5.187 | 1 | 4 | 16 |
| GO:0014911 | positive regulation of smooth muscle cell migration | 2.71e-02 | 1.00e+00 | 5.187 | 1 | 1 | 16 |
| GO:2000811 | negative regulation of anoikis | 2.71e-02 | 1.00e+00 | 5.187 | 1 | 1 | 16 |
| GO:0048037 | cofactor binding | 2.71e-02 | 1.00e+00 | 5.187 | 1 | 1 | 16 |
| GO:0005099 | Ras GTPase activator activity | 2.71e-02 | 1.00e+00 | 5.187 | 1 | 1 | 16 |
| GO:0042981 | regulation of apoptotic process | 2.75e-02 | 1.00e+00 | 2.949 | 2 | 7 | 151 |
| GO:0030742 | GTP-dependent protein binding | 2.88e-02 | 1.00e+00 | 5.100 | 1 | 1 | 17 |
| GO:0008284 | positive regulation of cell proliferation | 2.89e-02 | 1.00e+00 | 2.157 | 3 | 7 | 392 |
| GO:0010008 | endosome membrane | 2.95e-02 | 1.00e+00 | 2.892 | 2 | 2 | 157 |
| GO:0000278 | mitotic cell cycle | 3.00e-02 | 1.00e+00 | 2.135 | 3 | 16 | 398 |
| GO:0070064 | proline-rich region binding | 3.04e-02 | 1.00e+00 | 5.017 | 1 | 1 | 18 |
| GO:0031954 | positive regulation of protein autophosphorylation | 3.04e-02 | 1.00e+00 | 5.017 | 1 | 1 | 18 |
| GO:0090316 | positive regulation of intracellular protein transport | 3.04e-02 | 1.00e+00 | 5.017 | 1 | 1 | 18 |
| GO:0007088 | regulation of mitosis | 3.21e-02 | 1.00e+00 | 4.939 | 1 | 1 | 19 |
| GO:0045453 | bone resorption | 3.21e-02 | 1.00e+00 | 4.939 | 1 | 1 | 19 |
| GO:0031667 | response to nutrient levels | 3.21e-02 | 1.00e+00 | 4.939 | 1 | 1 | 19 |
| GO:0034220 | ion transmembrane transport | 3.31e-02 | 1.00e+00 | 2.803 | 2 | 2 | 167 |
| GO:0032148 | activation of protein kinase B activity | 3.38e-02 | 1.00e+00 | 4.865 | 1 | 1 | 20 |
| GO:0051090 | regulation of sequence-specific DNA binding transcription factor activity | 3.38e-02 | 1.00e+00 | 4.865 | 1 | 2 | 20 |
| GO:0043473 | pigmentation | 3.38e-02 | 1.00e+00 | 4.865 | 1 | 2 | 20 |
| GO:0072332 | intrinsic apoptotic signaling pathway by p53 class mediator | 3.38e-02 | 1.00e+00 | 4.865 | 1 | 1 | 20 |
| GO:0043393 | regulation of protein binding | 3.38e-02 | 1.00e+00 | 4.865 | 1 | 2 | 20 |
| GO:0046847 | filopodium assembly | 3.54e-02 | 1.00e+00 | 4.795 | 1 | 1 | 21 |
| GO:0007369 | gastrulation | 3.54e-02 | 1.00e+00 | 4.795 | 1 | 1 | 21 |
| GO:0051881 | regulation of mitochondrial membrane potential | 3.54e-02 | 1.00e+00 | 4.795 | 1 | 1 | 21 |
| GO:0071364 | cellular response to epidermal growth factor stimulus | 3.54e-02 | 1.00e+00 | 4.795 | 1 | 1 | 21 |
| GO:0007049 | cell cycle | 3.68e-02 | 1.00e+00 | 2.719 | 2 | 4 | 177 |
| GO:0001106 | RNA polymerase II transcription corepressor activity | 3.71e-02 | 1.00e+00 | 4.728 | 1 | 1 | 22 |
| GO:0031435 | mitogen-activated protein kinase kinase kinase binding | 3.71e-02 | 1.00e+00 | 4.728 | 1 | 1 | 22 |
| GO:2001243 | negative regulation of intrinsic apoptotic signaling pathway | 3.71e-02 | 1.00e+00 | 4.728 | 1 | 1 | 22 |
| GO:0046686 | response to cadmium ion | 3.71e-02 | 1.00e+00 | 4.728 | 1 | 1 | 22 |
| GO:0031625 | ubiquitin protein ligase binding | 3.79e-02 | 1.00e+00 | 2.695 | 2 | 4 | 180 |
| GO:0001892 | embryonic placenta development | 3.87e-02 | 1.00e+00 | 4.664 | 1 | 1 | 23 |
| GO:1900026 | positive regulation of substrate adhesion-dependent cell spreading | 3.87e-02 | 1.00e+00 | 4.664 | 1 | 1 | 23 |
| GO:0051297 | centrosome organization | 3.87e-02 | 1.00e+00 | 4.664 | 1 | 2 | 23 |
| GO:0045787 | positive regulation of cell cycle | 3.87e-02 | 1.00e+00 | 4.664 | 1 | 2 | 23 |
| GO:0002040 | sprouting angiogenesis | 3.87e-02 | 1.00e+00 | 4.664 | 1 | 1 | 23 |
| GO:0051017 | actin filament bundle assembly | 3.87e-02 | 1.00e+00 | 4.664 | 1 | 1 | 23 |
| GO:0015629 | actin cytoskeleton | 3.91e-02 | 1.00e+00 | 2.671 | 2 | 3 | 183 |
| GO:0006367 | transcription initiation from RNA polymerase II promoter | 3.94e-02 | 1.00e+00 | 2.664 | 2 | 5 | 184 |
| GO:0051602 | response to electrical stimulus | 4.04e-02 | 1.00e+00 | 4.602 | 1 | 1 | 24 |
| GO:0050715 | positive regulation of cytokine secretion | 4.20e-02 | 1.00e+00 | 4.543 | 1 | 1 | 25 |
| GO:0031234 | extrinsic component of cytoplasmic side of plasma membrane | 4.20e-02 | 1.00e+00 | 4.543 | 1 | 1 | 25 |
| GO:0048705 | skeletal system morphogenesis | 4.20e-02 | 1.00e+00 | 4.543 | 1 | 1 | 25 |
| GO:1900740 | positive regulation of protein insertion into mitochondrial membrane involved in apoptotic signaling pathway | 4.37e-02 | 1.00e+00 | 4.487 | 1 | 1 | 26 |
| GO:0071902 | positive regulation of protein serine/threonine kinase activity | 4.37e-02 | 1.00e+00 | 4.487 | 1 | 1 | 26 |
| GO:0045859 | regulation of protein kinase activity | 4.37e-02 | 1.00e+00 | 4.487 | 1 | 1 | 26 |
| GO:0031424 | keratinization | 4.53e-02 | 1.00e+00 | 4.432 | 1 | 1 | 27 |
| GO:0051149 | positive regulation of muscle cell differentiation | 4.53e-02 | 1.00e+00 | 4.432 | 1 | 2 | 27 |
| GO:0032720 | negative regulation of tumor necrosis factor production | 4.53e-02 | 1.00e+00 | 4.432 | 1 | 1 | 27 |
| GO:0001103 | RNA polymerase II repressing transcription factor binding | 4.53e-02 | 1.00e+00 | 4.432 | 1 | 1 | 27 |
| GO:0030331 | estrogen receptor binding | 4.53e-02 | 1.00e+00 | 4.432 | 1 | 1 | 27 |
| GO:2001238 | positive regulation of extrinsic apoptotic signaling pathway | 4.53e-02 | 1.00e+00 | 4.432 | 1 | 1 | 27 |
| GO:0031069 | hair follicle morphogenesis | 4.53e-02 | 1.00e+00 | 4.432 | 1 | 1 | 27 |
| GO:0032467 | positive regulation of cytokinesis | 4.70e-02 | 1.00e+00 | 4.380 | 1 | 1 | 28 |
| GO:0045944 | positive regulation of transcription from RNA polymerase II promoter | 4.80e-02 | 1.00e+00 | 1.524 | 4 | 11 | 811 |
| GO:0031252 | cell leading edge | 4.86e-02 | 1.00e+00 | 4.329 | 1 | 3 | 29 |
| GO:0048365 | Rac GTPase binding | 4.86e-02 | 1.00e+00 | 4.329 | 1 | 1 | 29 |
| GO:0034605 | cellular response to heat | 4.86e-02 | 1.00e+00 | 4.329 | 1 | 1 | 29 |
| GO:0072686 | mitotic spindle | 4.86e-02 | 1.00e+00 | 4.329 | 1 | 1 | 29 |
| GO:0042169 | SH2 domain binding | 5.03e-02 | 1.00e+00 | 4.280 | 1 | 1 | 30 |
| GO:0031647 | regulation of protein stability | 5.03e-02 | 1.00e+00 | 4.280 | 1 | 1 | 30 |
| GO:0005547 | phosphatidylinositol-3,4,5-trisphosphate binding | 5.03e-02 | 1.00e+00 | 4.280 | 1 | 1 | 30 |
| GO:0040018 | positive regulation of multicellular organism growth | 5.03e-02 | 1.00e+00 | 4.280 | 1 | 1 | 30 |
| GO:0046875 | ephrin receptor binding | 5.03e-02 | 1.00e+00 | 4.280 | 1 | 2 | 30 |
| GO:0070555 | response to interleukin-1 | 5.19e-02 | 1.00e+00 | 4.233 | 1 | 1 | 31 |
| GO:0007093 | mitotic cell cycle checkpoint | 5.19e-02 | 1.00e+00 | 4.233 | 1 | 2 | 31 |
| GO:0032320 | positive regulation of Ras GTPase activity | 5.35e-02 | 1.00e+00 | 4.187 | 1 | 1 | 32 |
| GO:0051219 | phosphoprotein binding | 5.35e-02 | 1.00e+00 | 4.187 | 1 | 4 | 32 |
| GO:0032091 | negative regulation of protein binding | 5.51e-02 | 1.00e+00 | 4.143 | 1 | 1 | 33 |
| GO:0033077 | T cell differentiation in thymus | 5.51e-02 | 1.00e+00 | 4.143 | 1 | 1 | 33 |
| GO:0005158 | insulin receptor binding | 5.51e-02 | 1.00e+00 | 4.143 | 1 | 2 | 33 |
| GO:0048812 | neuron projection morphogenesis | 5.51e-02 | 1.00e+00 | 4.143 | 1 | 1 | 33 |
| GO:0042692 | muscle cell differentiation | 5.68e-02 | 1.00e+00 | 4.100 | 1 | 2 | 34 |
| GO:0001890 | placenta development | 5.68e-02 | 1.00e+00 | 4.100 | 1 | 1 | 34 |
| GO:2001237 | negative regulation of extrinsic apoptotic signaling pathway | 5.84e-02 | 1.00e+00 | 4.058 | 1 | 1 | 35 |
| GO:0019221 | cytokine-mediated signaling pathway | 5.88e-02 | 1.00e+00 | 2.342 | 2 | 5 | 230 |
| GO:0007067 | mitotic nuclear division | 5.93e-02 | 1.00e+00 | 2.335 | 2 | 7 | 231 |
| GO:0071560 | cellular response to transforming growth factor beta stimulus | 6.00e-02 | 1.00e+00 | 4.017 | 1 | 1 | 36 |
| GO:0030178 | negative regulation of Wnt signaling pathway | 6.00e-02 | 1.00e+00 | 4.017 | 1 | 1 | 36 |
| GO:0016301 | kinase activity | 6.16e-02 | 1.00e+00 | 3.978 | 1 | 2 | 37 |
| GO:0018107 | peptidyl-threonine phosphorylation | 6.16e-02 | 1.00e+00 | 3.978 | 1 | 1 | 37 |
| GO:0032880 | regulation of protein localization | 6.16e-02 | 1.00e+00 | 3.978 | 1 | 1 | 37 |
| GO:0071277 | cellular response to calcium ion | 6.16e-02 | 1.00e+00 | 3.978 | 1 | 1 | 37 |
| GO:0097191 | extrinsic apoptotic signaling pathway | 6.32e-02 | 1.00e+00 | 3.939 | 1 | 3 | 38 |
| GO:0045740 | positive regulation of DNA replication | 6.32e-02 | 1.00e+00 | 3.939 | 1 | 1 | 38 |
| GO:0042542 | response to hydrogen peroxide | 6.65e-02 | 1.00e+00 | 3.865 | 1 | 1 | 40 |
| GO:0045785 | positive regulation of cell adhesion | 6.81e-02 | 1.00e+00 | 3.830 | 1 | 1 | 41 |
| GO:0070301 | cellular response to hydrogen peroxide | 6.81e-02 | 1.00e+00 | 3.830 | 1 | 1 | 41 |
| GO:0005902 | microvillus | 6.97e-02 | 1.00e+00 | 3.795 | 1 | 1 | 42 |
| GO:0004715 | non-membrane spanning protein tyrosine kinase activity | 6.97e-02 | 1.00e+00 | 3.795 | 1 | 1 | 42 |
| GO:0043025 | neuronal cell body | 7.00e-02 | 1.00e+00 | 2.198 | 2 | 5 | 254 |
| GO:0034613 | cellular protein localization | 7.29e-02 | 1.00e+00 | 3.728 | 1 | 2 | 44 |
| GO:0005080 | protein kinase C binding | 7.29e-02 | 1.00e+00 | 3.728 | 1 | 1 | 44 |
| GO:0006915 | apoptotic process | 7.31e-02 | 1.00e+00 | 1.615 | 3 | 9 | 571 |
| GO:0045860 | positive regulation of protein kinase activity | 7.45e-02 | 1.00e+00 | 3.695 | 1 | 1 | 45 |
| GO:0009411 | response to UV | 7.45e-02 | 1.00e+00 | 3.695 | 1 | 2 | 45 |
| GO:0006281 | DNA repair | 7.48e-02 | 1.00e+00 | 2.143 | 2 | 5 | 264 |
| GO:0021762 | substantia nigra development | 7.60e-02 | 1.00e+00 | 3.664 | 1 | 2 | 46 |
| GO:0016328 | lateral plasma membrane | 7.60e-02 | 1.00e+00 | 3.664 | 1 | 3 | 46 |
| GO:0044297 | cell body | 7.60e-02 | 1.00e+00 | 3.664 | 1 | 1 | 46 |
| GO:0043525 | positive regulation of neuron apoptotic process | 7.60e-02 | 1.00e+00 | 3.664 | 1 | 2 | 46 |
| GO:0045727 | positive regulation of translation | 7.60e-02 | 1.00e+00 | 3.664 | 1 | 1 | 46 |
| GO:0005884 | actin filament | 7.60e-02 | 1.00e+00 | 3.664 | 1 | 2 | 46 |
| GO:0043406 | positive regulation of MAP kinase activity | 7.76e-02 | 1.00e+00 | 3.632 | 1 | 1 | 47 |
| GO:0008344 | adult locomotory behavior | 7.76e-02 | 1.00e+00 | 3.632 | 1 | 2 | 47 |
| GO:0006950 | response to stress | 7.92e-02 | 1.00e+00 | 3.602 | 1 | 1 | 48 |
| GO:0005975 | carbohydrate metabolic process | 7.98e-02 | 1.00e+00 | 2.089 | 2 | 3 | 274 |
| GO:0005070 | SH3/SH2 adaptor activity | 8.08e-02 | 1.00e+00 | 3.572 | 1 | 1 | 49 |
| GO:0007030 | Golgi organization | 8.24e-02 | 1.00e+00 | 3.543 | 1 | 3 | 50 |
| GO:0090263 | positive regulation of canonical Wnt signaling pathway | 8.40e-02 | 1.00e+00 | 3.515 | 1 | 1 | 51 |
| GO:0030900 | forebrain development | 8.40e-02 | 1.00e+00 | 3.515 | 1 | 1 | 51 |
| GO:0003684 | damaged DNA binding | 8.40e-02 | 1.00e+00 | 3.515 | 1 | 2 | 51 |
| GO:0045732 | positive regulation of protein catabolic process | 8.40e-02 | 1.00e+00 | 3.515 | 1 | 1 | 51 |
| GO:0008202 | steroid metabolic process | 8.55e-02 | 1.00e+00 | 3.487 | 1 | 1 | 52 |
| GO:0019899 | enzyme binding | 8.69e-02 | 1.00e+00 | 2.017 | 2 | 5 | 288 |
| GO:0006952 | defense response | 8.71e-02 | 1.00e+00 | 3.459 | 1 | 1 | 53 |
| GO:0030175 | filopodium | 8.71e-02 | 1.00e+00 | 3.459 | 1 | 2 | 53 |
| GO:0007264 | small GTPase mediated signal transduction | 8.79e-02 | 1.00e+00 | 2.007 | 2 | 7 | 290 |
| GO:0009612 | response to mechanical stimulus | 8.87e-02 | 1.00e+00 | 3.432 | 1 | 1 | 54 |
| GO:0097193 | intrinsic apoptotic signaling pathway | 9.03e-02 | 1.00e+00 | 3.406 | 1 | 2 | 55 |
| GO:0002039 | p53 binding | 9.03e-02 | 1.00e+00 | 3.406 | 1 | 1 | 55 |
| GO:0046330 | positive regulation of JNK cascade | 9.03e-02 | 1.00e+00 | 3.406 | 1 | 1 | 55 |
| GO:0043234 | protein complex | 9.31e-02 | 1.00e+00 | 1.958 | 2 | 9 | 300 |
| GO:0005793 | endoplasmic reticulum-Golgi intermediate compartment | 9.34e-02 | 1.00e+00 | 3.354 | 1 | 1 | 57 |
| GO:0005886 | plasma membrane | 9.85e-02 | 1.00e+00 | 0.718 | 8 | 24 | 2834 |
| GO:0005794 | Golgi apparatus | 9.88e-02 | 1.00e+00 | 1.428 | 3 | 8 | 650 |
| GO:0033138 | positive regulation of peptidyl-serine phosphorylation | 9.96e-02 | 1.00e+00 | 3.256 | 1 | 1 | 61 |
| GO:0005739 | mitochondrion | 1.01e-01 | 1.00e+00 | 1.156 | 4 | 10 | 1046 |
| GO:0001078 | RNA polymerase II core promoter proximal region sequence-specific DNA binding transcription factor activity involved in negative regulation of transcription | 1.01e-01 | 1.00e+00 | 3.233 | 1 | 1 | 62 |
| GO:0000151 | ubiquitin ligase complex | 1.03e-01 | 1.00e+00 | 3.210 | 1 | 1 | 63 |
| GO:0005901 | caveola | 1.03e-01 | 1.00e+00 | 3.210 | 1 | 1 | 63 |
| GO:0019903 | protein phosphatase binding | 1.03e-01 | 1.00e+00 | 3.210 | 1 | 1 | 63 |
| GO:0042995 | cell projection | 1.03e-01 | 1.00e+00 | 3.210 | 1 | 1 | 63 |
| GO:0043154 | negative regulation of cysteine-type endopeptidase activity involved in apoptotic process | 1.03e-01 | 1.00e+00 | 3.210 | 1 | 2 | 63 |
| GO:0006417 | regulation of translation | 1.03e-01 | 1.00e+00 | 3.210 | 1 | 4 | 63 |
| GO:0032869 | cellular response to insulin stimulus | 1.04e-01 | 1.00e+00 | 3.187 | 1 | 2 | 64 |
| GO:0016491 | oxidoreductase activity | 1.06e-01 | 1.00e+00 | 3.165 | 1 | 1 | 65 |
| GO:0071260 | cellular response to mechanical stimulus | 1.07e-01 | 1.00e+00 | 3.143 | 1 | 3 | 66 |
| GO:0007411 | axon guidance | 1.08e-01 | 1.00e+00 | 1.834 | 2 | 3 | 327 |
| GO:0030141 | secretory granule | 1.09e-01 | 1.00e+00 | 3.121 | 1 | 2 | 67 |
| GO:0043231 | intracellular membrane-bounded organelle | 1.10e-01 | 1.00e+00 | 1.812 | 2 | 3 | 332 |
| GO:0035264 | multicellular organism growth | 1.13e-01 | 1.00e+00 | 3.058 | 1 | 1 | 70 |
| GO:0042393 | histone binding | 1.15e-01 | 1.00e+00 | 3.037 | 1 | 1 | 71 |
| GO:0001503 | ossification | 1.15e-01 | 1.00e+00 | 3.037 | 1 | 2 | 71 |
| GO:0043086 | negative regulation of catalytic activity | 1.20e-01 | 1.00e+00 | 2.978 | 1 | 1 | 74 |
| GO:0042826 | histone deacetylase binding | 1.20e-01 | 1.00e+00 | 2.978 | 1 | 1 | 74 |
| GO:0060070 | canonical Wnt signaling pathway | 1.21e-01 | 1.00e+00 | 2.958 | 1 | 2 | 75 |
| GO:0007265 | Ras protein signal transduction | 1.21e-01 | 1.00e+00 | 2.958 | 1 | 3 | 75 |
| GO:0051897 | positive regulation of protein kinase B signaling | 1.21e-01 | 1.00e+00 | 2.958 | 1 | 1 | 75 |
| GO:0044325 | ion channel binding | 1.24e-01 | 1.00e+00 | 2.920 | 1 | 3 | 77 |
| GO:0007229 | integrin-mediated signaling pathway | 1.26e-01 | 1.00e+00 | 2.902 | 1 | 2 | 78 |
| GO:0071013 | catalytic step 2 spliceosome | 1.27e-01 | 1.00e+00 | 2.883 | 1 | 1 | 79 |
| GO:0010629 | negative regulation of gene expression | 1.29e-01 | 1.00e+00 | 2.865 | 1 | 2 | 80 |
| GO:0006351 | transcription, DNA-templated | 1.30e-01 | 1.00e+00 | 0.879 | 5 | 17 | 1585 |
| GO:0050796 | regulation of insulin secretion | 1.32e-01 | 1.00e+00 | 2.830 | 1 | 1 | 82 |
| GO:0045177 | apical part of cell | 1.32e-01 | 1.00e+00 | 2.830 | 1 | 1 | 82 |
| GO:0004713 | protein tyrosine kinase activity | 1.32e-01 | 1.00e+00 | 2.830 | 1 | 1 | 82 |
| GO:0030336 | negative regulation of cell migration | 1.33e-01 | 1.00e+00 | 2.812 | 1 | 2 | 83 |
| GO:0005179 | hormone activity | 1.35e-01 | 1.00e+00 | 2.795 | 1 | 1 | 84 |
| GO:0042593 | glucose homeostasis | 1.42e-01 | 1.00e+00 | 2.711 | 1 | 1 | 89 |
| GO:0042384 | cilium assembly | 1.44e-01 | 1.00e+00 | 2.695 | 1 | 2 | 90 |
| GO:0000922 | spindle pole | 1.45e-01 | 1.00e+00 | 2.679 | 1 | 5 | 91 |
| GO:0016337 | single organismal cell-cell adhesion | 1.47e-01 | 1.00e+00 | 2.664 | 1 | 2 | 92 |
| GO:0006928 | cellular component movement | 1.47e-01 | 1.00e+00 | 2.664 | 1 | 4 | 92 |
| GO:0005200 | structural constituent of cytoskeleton | 1.48e-01 | 1.00e+00 | 2.648 | 1 | 6 | 93 |
| GO:0005770 | late endosome | 1.49e-01 | 1.00e+00 | 2.632 | 1 | 1 | 94 |
| GO:0030426 | growth cone | 1.54e-01 | 1.00e+00 | 2.587 | 1 | 3 | 97 |
| GO:0000139 | Golgi membrane | 1.54e-01 | 1.00e+00 | 1.515 | 2 | 7 | 408 |
| GO:0051015 | actin filament binding | 1.54e-01 | 1.00e+00 | 2.587 | 1 | 1 | 97 |
| GO:0005178 | integrin binding | 1.54e-01 | 1.00e+00 | 2.587 | 1 | 1 | 97 |
| GO:0006112 | energy reserve metabolic process | 1.57e-01 | 1.00e+00 | 2.558 | 1 | 2 | 99 |
| GO:0014069 | postsynaptic density | 1.67e-01 | 1.00e+00 | 2.459 | 1 | 3 | 106 |
| GO:0005096 | GTPase activator activity | 1.70e-01 | 1.00e+00 | 2.432 | 1 | 3 | 108 |
| GO:0005815 | microtubule organizing center | 1.73e-01 | 1.00e+00 | 2.406 | 1 | 4 | 110 |
| GO:0070374 | positive regulation of ERK1 and ERK2 cascade | 1.73e-01 | 1.00e+00 | 2.406 | 1 | 1 | 110 |
| GO:0020037 | heme binding | 1.74e-01 | 1.00e+00 | 2.393 | 1 | 1 | 111 |
| GO:0050900 | leukocyte migration | 1.74e-01 | 1.00e+00 | 2.393 | 1 | 1 | 111 |
| GO:0007596 | blood coagulation | 1.89e-01 | 1.00e+00 | 1.329 | 2 | 5 | 464 |
| GO:0032496 | response to lipopolysaccharide | 1.90e-01 | 1.00e+00 | 2.256 | 1 | 1 | 122 |
| GO:0051056 | regulation of small GTPase mediated signal transduction | 1.91e-01 | 1.00e+00 | 2.245 | 1 | 3 | 123 |
| GO:0030036 | actin cytoskeleton organization | 1.99e-01 | 1.00e+00 | 2.176 | 1 | 3 | 129 |
| GO:0046983 | protein dimerization activity | 2.02e-01 | 1.00e+00 | 2.154 | 1 | 3 | 131 |
| GO:0045893 | positive regulation of transcription, DNA-templated | 2.03e-01 | 1.00e+00 | 1.259 | 2 | 8 | 487 |
| GO:0018108 | peptidyl-tyrosine phosphorylation | 2.05e-01 | 1.00e+00 | 2.132 | 1 | 1 | 133 |
| GO:0016055 | Wnt signaling pathway | 2.16e-01 | 1.00e+00 | 2.048 | 1 | 3 | 141 |
| GO:0005911 | cell-cell junction | 2.17e-01 | 1.00e+00 | 2.037 | 1 | 2 | 142 |
| GO:0008286 | insulin receptor signaling pathway | 2.20e-01 | 1.00e+00 | 2.017 | 1 | 4 | 144 |
| GO:0055085 | transmembrane transport | 2.20e-01 | 1.00e+00 | 1.181 | 2 | 3 | 514 |
| GO:0007165 | signal transduction | 2.21e-01 | 1.00e+00 | 0.880 | 3 | 7 | 950 |
| GO:0046777 | protein autophosphorylation | 2.39e-01 | 1.00e+00 | 1.883 | 1 | 1 | 158 |
| GO:0008022 | protein C-terminus binding | 2.43e-01 | 1.00e+00 | 1.856 | 1 | 4 | 161 |
| GO:0005516 | calmodulin binding | 2.44e-01 | 1.00e+00 | 1.847 | 1 | 3 | 162 |
| GO:0000398 | mRNA splicing, via spliceosome | 2.48e-01 | 1.00e+00 | 1.821 | 1 | 2 | 165 |
| GO:0000978 | RNA polymerase II core promoter proximal region sequence-specific DNA binding | 2.52e-01 | 1.00e+00 | 1.795 | 1 | 4 | 168 |
| GO:0030424 | axon | 2.57e-01 | 1.00e+00 | 1.761 | 1 | 4 | 172 |
| GO:0016607 | nuclear speck | 2.61e-01 | 1.00e+00 | 1.736 | 1 | 2 | 175 |
| GO:0004672 | protein kinase activity | 2.65e-01 | 1.00e+00 | 1.711 | 1 | 4 | 178 |
| GO:0005764 | lysosome | 2.70e-01 | 1.00e+00 | 1.679 | 1 | 2 | 182 |
| GO:0032403 | protein complex binding | 2.73e-01 | 1.00e+00 | 1.656 | 1 | 2 | 185 |
| GO:0043547 | positive regulation of GTPase activity | 2.80e-01 | 1.00e+00 | 1.617 | 1 | 4 | 190 |
| GO:0005654 | nucleoplasm | 2.89e-01 | 1.00e+00 | 0.675 | 3 | 26 | 1095 |
| GO:0003924 | GTPase activity | 2.96e-01 | 1.00e+00 | 1.522 | 1 | 6 | 203 |
| GO:0030168 | platelet activation | 2.98e-01 | 1.00e+00 | 1.508 | 1 | 4 | 205 |
| GO:0004871 | signal transducer activity | 3.12e-01 | 1.00e+00 | 1.432 | 1 | 1 | 216 |
| GO:0006184 | GTP catabolic process | 3.16e-01 | 1.00e+00 | 1.406 | 1 | 6 | 220 |
| GO:0016874 | ligase activity | 3.21e-01 | 1.00e+00 | 1.380 | 1 | 2 | 224 |
| GO:0030425 | dendrite | 3.40e-01 | 1.00e+00 | 1.280 | 1 | 3 | 240 |
| GO:0004842 | ubiquitin-protein transferase activity | 3.58e-01 | 1.00e+00 | 1.187 | 1 | 3 | 256 |
| GO:0005874 | microtubule | 3.60e-01 | 1.00e+00 | 1.176 | 1 | 3 | 258 |
| GO:0005102 | receptor binding | 3.71e-01 | 1.00e+00 | 1.121 | 1 | 2 | 268 |
| GO:0000166 | nucleotide binding | 3.76e-01 | 1.00e+00 | 1.100 | 1 | 2 | 272 |
| GO:0007283 | spermatogenesis | 3.80e-01 | 1.00e+00 | 1.079 | 1 | 2 | 276 |
| GO:0044281 | small molecule metabolic process | 3.85e-01 | 1.00e+00 | 0.433 | 3 | 16 | 1295 |
| GO:0042493 | response to drug | 3.93e-01 | 1.00e+00 | 1.017 | 1 | 2 | 288 |
| GO:0005743 | mitochondrial inner membrane | 4.05e-01 | 1.00e+00 | 0.958 | 1 | 1 | 300 |
| GO:0005524 | ATP binding | 4.14e-01 | 1.00e+00 | 0.369 | 3 | 19 | 1354 |
| GO:0004674 | protein serine/threonine kinase activity | 4.18e-01 | 1.00e+00 | 0.902 | 1 | 6 | 312 |
| GO:0035556 | intracellular signal transduction | 4.23e-01 | 1.00e+00 | 0.879 | 1 | 5 | 317 |
| GO:0005525 | GTP binding | 4.34e-01 | 1.00e+00 | 0.830 | 1 | 6 | 328 |
| GO:0008283 | cell proliferation | 4.37e-01 | 1.00e+00 | 0.816 | 1 | 4 | 331 |
| GO:0005813 | centrosome | 4.45e-01 | 1.00e+00 | 0.782 | 1 | 9 | 339 |
| GO:0007275 | multicellular organismal development | 4.50e-01 | 1.00e+00 | 0.761 | 1 | 2 | 344 |
| GO:0003723 | RNA binding | 4.60e-01 | 1.00e+00 | 0.715 | 1 | 10 | 355 |
| GO:0030054 | cell junction | 4.61e-01 | 1.00e+00 | 0.711 | 1 | 4 | 356 |
| GO:0008285 | negative regulation of cell proliferation | 4.71e-01 | 1.00e+00 | 0.667 | 1 | 3 | 367 |
| GO:0045892 | negative regulation of transcription, DNA-templated | 5.22e-01 | 1.00e+00 | 0.459 | 1 | 2 | 424 |
| GO:0006366 | transcription from RNA polymerase II promoter | 5.23e-01 | 1.00e+00 | 0.456 | 1 | 3 | 425 |
| GO:0006468 | protein phosphorylation | 5.57e-01 | 1.00e+00 | 0.320 | 1 | 6 | 467 |
| GO:0055114 | oxidation-reduction process | 5.68e-01 | 1.00e+00 | 0.277 | 1 | 2 | 481 |
| GO:0006355 | regulation of transcription, DNA-templated | 5.74e-01 | 1.00e+00 | 0.079 | 2 | 10 | 1104 |
| GO:0042802 | identical protein binding | 5.75e-01 | 1.00e+00 | 0.248 | 1 | 4 | 491 |
| GO:0048471 | perinuclear region of cytoplasm | 5.99e-01 | 1.00e+00 | 0.156 | 1 | 9 | 523 |
| GO:0005509 | calcium ion binding | 6.43e-01 | 1.00e+00 | -0.015 | 1 | 4 | 589 |
| GO:0005783 | endoplasmic reticulum | 6.56e-01 | 1.00e+00 | -0.066 | 1 | 6 | 610 |
| GO:0003700 | sequence-specific DNA binding transcription factor activity | 7.31e-01 | 1.00e+00 | -0.360 | 1 | 9 | 748 |
| GO:0005615 | extracellular space | 8.33e-01 | 1.00e+00 | -0.793 | 1 | 3 | 1010 |
| GO:0008270 | zinc ion binding | 8.50e-01 | 1.00e+00 | -0.872 | 1 | 7 | 1067 |
| GO:0003677 | DNA binding | 9.11e-01 | 1.00e+00 | -1.213 | 1 | 14 | 1351 |
| GO:0046872 | metal ion binding | 9.28e-01 | 1.00e+00 | -1.330 | 1 | 14 | 1465 |